Entries are categorized by year. Within each year, they are listed alphabetically. It should be noted that the dates of some entries may vary depending on what is considered to mark the specific event (e.g. the date when the event appeared in a publication, the date when a company was founded, the date when a company was made public, etc.) As such, this list should act as a loose timeline of progress in neuroengineering fields rather than a temporally exact record. I hope that you enjoy reading about these exciting breakthroughs in neurotechnology, connectomics, and brain simulation!
PDF version: Global Highlights in Neurotechnology, Connectomics, and Brain Simulation 2005 to 2019 – by Logan Thrasher Collins
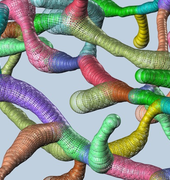
Cover image source: Motta et al., 2019
2005
1 – Optogenetic stimulation using ChR2
Reference: (Boyden, Zhang, Bamberg, Nagel, & Deisseroth, 2005)
Neurons were engineered to express a bacterial light-gated ion channel called ChR2 which responds to blue wavelengths. In this way, the neurons were stimulated to fire using blue light. This innovation gave rise to the field of optogenetics.

2006
2 – Blue Brain Project and detailed cortical column simulation
Reference: (Markram, 2006)
Henry Markram announced the European Blue Brain Project (which later developed into the Human Brain Project), a large-scale initiative with the goal of simulating the brain in a supercomputer. In November 2007, the Blue Brain Project reported a biophysically detailed simulation of a rat neocortical column consisting of about 10,000multicompartmental Hodgkin-Huxley neuron models and over ten million synapses. The connectivity was based on the numbers of inputs and outputs quantified for given cell types. The simulation was carried out using the Blue Gene/L supercomputer.

2007
3 – Brainbow
Reference: (Livet et al., 2007)
Neurons were genetically engineered using Cre/Lox tools to randomly express three or more differently-colored fluorescent proteins (XFPs) in a given cell, labeling that cell with a unique combination of colors. Ninety distinct colors were emitted from a population of these engineered neurons. Tissue modified with Brainbow allows superior imaging since neurons can be distinguished via color contrast.

4 – Optogenetic silencing using halorhodopsin
Reference: (Han & Boyden, 2007)
A bacterial protein channel called halorhodopsin was expressed in neurons (along with ChR2). Halorhodopsin moves chloride ions into the cell upon exposure to yellow light, hyperpolarizing the membrane and inhibiting neuronal activity. By using both halorhodopsin and ChR2, neurons were inhibited and activated with yellow and blue light respectively.

2008
5 – Anatomically simplified simulation of 22 million neurons
Reference: (Djurfeldt et al., 2008)
As part of the European FACETs project, the IBM Blue Gene/L supercomputer was employed to carry out a simulation consisting of 22 million simplified multicompartmental Hodgkin-Huxley neurons with 11 billion synapses. Depending on the cell type, the neurons were modeled using 3-6 compartments. Pyramidal cells and two types of inhibitory interneuron were used. Connectivity was defined according to a simple set of rules which acted to roughly reflect biological trends. With this setup, subpopulations of virtual neurons were able to store memories in the form of activity patterns.

2010
6 – Hardware for faster than real-time neuroscience simulations
Reference: (Schemmel et al., 2010)
As part of the European FACETs project, the neuromorphic hardware system BrainScaleS was developed to simulate networks of adaptive exponential integrate-and-fire neuron models 1,000-100,000 times faster than biological real-time. With this system, each neuromorphic wafer can simulate up to 180,000 neurons and forty million synapses and each artificial neuron can receive up to 14,336 presynaptic inputs. Multiple wafers can be interconnected to simulate larger networks. In addition, a Python-based software interface called PyNN was developed to facilitate easier configuration of the wafers to simulate desired neuronal networks.

7 – High temporal precision optogenetics
Reference: (Gunaydin et al., 2010)
The ChR2 protein was mutated to make a new optogenetic ion channel dubbed ChETA, allowing light-mediated induction of spike trains with much higher frequencies (up to 200 Hz) and decreasing the incidence of unintended spikes. ChETA also eliminates a phenomenon which can interfere with precise control of neural activity called the plateau potential.

2011
8 – Automated tape collecting ultramicrotome
Reference: (Schalek et al., 2011)
To increase the throughput of scanning electron microscopy (SEM) for imaging brain tissue, the automated tape collecting ultramicrotome (ATUM) was developed. ATUM automatically slices tissue volumes into thin sections (around 30 nm thickness) and deposits the sections onto a continuous tape. Many pieces of the tape can easily be put onto silicon wafers, stained, and automatically imaged via SEM. After imaging, the wafers can be stored for later use. Though this method increases the throughput of the SEM process, ATUM-SEM alone still can take weeks or months to image volumes of just a few hundred micrometers on a side. Nonetheless, ATUM-SEM has paved the way for more advances in volume SEM.

2012
9 – Hippocampal prosthesis in rats
Reference: (Berger et al., 2012)
An electronic prosthesis was shown to restore memory to rats with pharmacologically inhibited hippocampal function as well as to enhance memory in rats with normally functioning hippocampi. This prosthesis consisted of recording electrodes at the CA3 region of the hippocampus, stimulation electrodes at the CA1 region of the hippocampus, and a mathematical signal processing model which took inputs from the recording electrodes and predicted an appropriate set of outputs to send via the stimulation electrodes.

10 – In vivo superresolution microscopy to visualize neurons
Reference: (Berning, Willig, Steffens, Dibaj, & Hell, 2012)
Stimulated emission depletion microscopy (STED), a type of superresolution microscopy, was used to observe neurons in live anesthetized mice (equipped with skull windows) at a resolution of <70 nm. Dendritic spine morphologies were visible up to 15 μm beneath the surface of the brain tissue. Images were taken every few minutes and so illustrated how the morphologies of the spines changed over time.

2013
11 – CLARITY method
Reference: (Chung & Deisseroth, 2013)
CLARITY was developed as a method to make neural tissue samples translucent without damaging cellular morphologies. Tissue was infused with hydrogel monomers which were then crosslinked. Lipids were extracted using organic solvents and electrophoresis. CLARITY allows immunohistochemistry and fluorescence microscopy to be performed on volumetric tissue samples (or even entire brains) as well as making the tissue more permeable to macromolecules such as antibodies.

12 – Eyewire as a crowdsourcing method for retina mapping
Reference: (Marx, 2013)
To facilitate reconstruction of neurons in the retina (from serial section electron microscopy data), Eyewire employs crowdsourcing in the form of an online game. Images of tissue slices are given to the players who then “color in” the parts of the images corresponding to cells. Reconstructions are made by stacking many of these images on top of each other. As of the end of 2019, over 700 neurons have been mapped by Eyewire players.

13 – In vivo three-photon microscopy
Reference: (Horton et al., 2013)
In multi-photon microscopy, more than one photon of lower energy recombines during the excitation of a fluorophore, causing the fluorophore to emit a higher energy photon. Since long wavelengths of excitation light are employed, deep tissue penetration is possible. Using a three-photon method with excitation wavelengths of 1,700 nm, imaging was performed at a depth of up to 1.4 mm in living mouse brains. Blood vessels and RFP-labeled neurons were imaged using this approach.

14 – Telepathic rats engineered using hippocampal prosthesis
Reference: (Deadwyler et al., 2013)
The hippocampal prosthesis from highlight #9 was implanted in pairs of rats. Donor rats were trained to perform a task and their prostheses recorded the patterns of neural activity. These patterns were transmitted to recipient rats which had not been trained on the task, allowing the recipient rats to perform significantly better at the task compared to control animals.

15 – The BRAIN Initiative
Reference: (“Fact Sheet: BRAIN Initiative,” 2013)
The BRAIN Initiative (Brain Research through Advancing Innovative Technologies) provided neuroscientists with $110 million in governmental funding and $122 million in funding from private sources such as HHMI and the Allen Institute. It emphasizes research towards new tools for functional decoding of neural circuits and encourages collaboration between neuroscientists and physicists.

16 – Whole-brain functional recording from larval zebrafish
Reference: (Ahrens, Orger, Robson, Li, & Keller, 2013)
Light-sheet microscopy, a genetically encoded calcium sensor, and computational analysis methods were used to image larval zebrafish brain activity at single-cell resolution from about 80% of the total neurons in their brains (larval zebrafish are translucent). Patterns of correlated activity were identified among different groups of neurons, showing the technique’s utility for uncovering neural circuit dynamics.

17 – X-ray microscopy reconstructs Drosophila hemisphere
Reference: (Mizutani, Saiga, Takeuchi, Uesugi, & Suzuki, 2013)
The Drosophila brain was stained with silver nitrate and tetrachloroaurate to allow 3D imaging via synchrotron-based x-ray microscopy at a voxel size of 220 × 328 × 314 nm. Manual tracing (about 1,700 person hours) was performed to segment an entire hemisphere of the Drosophila brain. Some neuronal filaments were erroneously merged or were fragmented and removed from consideration. As a result, the final reconstruction only included about a third of the hemisphere’s total estimated number of traces. Nonetheless, this study set a foundation for future connectomics research with improved technologies.

2014
18 – Fluorescent in situ sequencing of RNAs
Reference: (Lee et al., 2014)
With fluorescent in situ sequencing (FISSEQ), RNAs undergo sequencing within intact tissue, allowing mapping of their spatial locations. Tissue is fixed on a glass slide, reverse transcriptase and CircLigase are used to convert the sample’s RNAs to circular cDNAs, the cDNAs are crosslinked into the tissue to keep them at the original locations of the RNAs, and the cDNAs are amplified using rolling circle amplification. Primers are then added and the tissue is washed with fluorophore-linked oligonucleotides which bind to specific dinucleotide pairs on the cDNAs. After each wash, the tissue is imaged and the colors of the fluorophores indicate the sequences of the cDNAs. To prevent confusion, the fluorophores are cleaved off after each cycle. Since the oligonucleotides are eight bases long (and they can only sequence two nucleotides at a time), this process is repeated with starting primers of different lengths to fill the gaps in the sequences. Novel software is used to facilitate image analysis to allow for accurate sequencing. Using FISSEQ, the spatial locations of RNAs (and ~20-30 base pair subsets of their sequences) representing thousands of unique genes can be found in tissue samples. FISSEQ has also been proposed as a connectomics strategy since it would allow unique barcoding of neurons and their synapses.

19 – Integrated Information Theory 3.0
Reference: (Oizumi, Albantakis, & Tononi, 2014)
Integrated information theory (IIT) is a quantitative theory of consciousness (originally proposed in 2004). IIT assumes some phenomenological axions and assumes causality to help define maximally irreducible conceptual structures (MICS) which represent conscious experiences which cannot be broken down into smaller parts. IIT also outlines a mathematical measure of an experience’s quantity called integrated information or ϕ.

20 – OpenWorm
Reference: (Szigeti et al., 2014)
The C. elegans connectome was mapped in 1986. Since then, more data on neurotransmitters, electrophysiology, cellular morphology, etc. has been collected. OpenWorm seeks to leverage these data towards creating a biologically realistic virtual worm. The project emphasizes collaboration, open access, and crowdsourcing. OpenWorm has developed software called Geppetto to facilitate integration of multicompartmental Hodgkin-Huxley models with soft-body physics simulations to help simulate the worm’s nervous system, musculature, and environment.

2015
21 – BigNeuron software for standardizing neuronal tracing
Reference: (Peng et al., 2015)
BigNeuron was established to counter a lack of standardization for neuronal reconstruction methods. Using high-performance computing, it tests numerous automated neuronal reconstruction algorithms on large-scale microscopy datasets. The algorithms are added as plugins to a central software platform called Vaa3D. BigNeuron aims to create a superior community-oriented morphological database and a set of more standardized tools for reconstruction.
22 – Detailed reconstruction of a small volume of mouse cortex
Reference: (Kasthuri et al., 2015)
Using automated tape collecting ultramicrotome scanning electron microscopy (ATUM-SEM), a 40×40×50 μm volume of neocortex was imaged at 3×3×30 nm voxel size. Within this volume, segmentation was performed on a volume of 1,500 μm3. Part of this volume underwent automatic segmentation before manual correction of errors and the rest was segmented manually. Synaptic connectivity, mitochondrial morphology, dendritic spine numbers and sizes, and synaptic vesicle locations and counts were quantified in parts of the volume. The connectivity patterns observed in this dataset indicated that synaptic connections are not predictable just from the proximity of axons and dendrites.

23 – Expansion microscopy
Reference: (F. Chen, Tillberg, & Boyden, 2015)
Expansion microscopy (ExM) is a process by which tissue can be physically enlarged in an isotropic fashion, increasing the effective resolution of any microscopy technique. The process involves infusion with a swellable polymer, crosslinking of biomolecules with the gel, polymerization, and water-mediated expansion. ExM typically enlarges tissue 4-fold and introduces only ~1% distortion. ExM greatly increases the optical translucency of treated tissue and makes the tissue more permeable to immunohistochemical stains.

24 – Human Brain Project cortical mesocircuit simulation
Reference: (Markram et al., 2015)
By integrating many different types of biological data (morphological cell types, electrical cell types, electrophysiological recordings, connectivity patterns, etc.), a virtual version of a 0.29 mm3 region of rat cortex with about 31,000 neurons and 37 million synapses was built. The simulation employed multicompartmental Hodgkin-Huxley neurons. Morphologies were determined by inserting many copies of 55 morphological neuron types at data-derived densities into each cortical layer. Connectivity was decided according to an algorithm which took into account the layer-specific densities of each neuron type, the lengths of axons for each neuron type, the densities of synaptic boutons coming from the axons of each neuron type, the mean number of synapses per connection between neurons for each pair of morphological neuron types, and the connection probability for each pair of morphological neuron types. This cortical mesocircuit was run on the Blue Gene/Q supercomputer. Activity similar to the biological equivalent emerged and new insights on neuronal computations were acquired.

25 – Human telepathy during a 20 questions game
Reference: (Stocco et al., 2015)
Telepathy between humans was achieved using electroencephalography (EEG) and transcranial magnetic stimulation (TMS). Pairs of people played 20 questions to identify unknown objects. EEG data from one person were recorded, processed, and transmitted to a second person via TMS occipital lobe stimulation. Higher-intensity TMS pulses corresponded to “yes” and lower-intensity TMS pulses corresponded to “no.” By comparison to controls, the brain-brain interface users were significantly more successful at playing 20 questions.

26 – Japan’s Brain/MINDS project
Reference: (Okano, Miyawaki, & Kasai, 2015)
Japan’s Brain/MINDS project aims to improve understanding of the brain using the marmoset as a model organism. It received almost $30 million in funding for its first year alone. The project is developing new tools for mapping brains and is seeking to find new treatments for brain disease.

27 – Multibeam SEM for connectomics
Reference: (Eberle et al., 2015)
Scanning electron microscopy (SEM) efficiency is severely limited by the amount of time the electron beam takes to image a single pixel. To increase the acquisition rate, an SEM device with a hexagonal array of 61 electron beams (instead of just one beam) was constructed. For imaging a hypothetical 1 mm3 volume, it was estimated that the acquisition would take only 6 months. This would be about 50 times faster than using a conventional SEM. Multibeam SEM was designed for compatibility with both serial section and block-face methods.

28 – Multibeam SEM with even more beams
Reference: (Kemen et al., 2015)
Due to the modularity of the multibeam scanning electron microscopy (SEM) design, a multibeam SEM with 91 beams and a corresponding larger number of detectors was built. In this multibeam SEM, a larger current per beam was achieved, granting better signal-to-noise ratios for imaging. The greater number of beams further increases the throughput of imaging relative to that of the 61 beam device. This version of the multibeam SEM also paves the way for more increases to the number of beams and consequent faster imaging.

29 – Neural lace
Reference: (Liu et al., 2015)
Neural lace consists of a syringe-injectable flexible electronic mesh made from submicrometer-thick wiring for neural interfacing. Upon injection, the mesh expands to record from centimeter-scale regions of tissue. Neural lace may enable less invasive brain-computer interfaces which circumvent the need for surgical implantation. Further development of this technology towards clinical application is underway.

30 – Recording from C. elegans neurons reveals motor operations
Reference: (Kato et al., 2015)
With the calcium indicator GCaMP, neural activity was recorded from all of the head ganglion neurons and some of the motor system neurons of live C. elegans. Since the C. elegans connectome has been characterized, most of these neurons were identifiable. Because more than 100 neurons were recorded, principal component analysis and phase space plots were employed to visualize the key patterns within the data. Using the phase space plots, specific motor actions were found to correspond with specific sequences of neural events.

31 – The TrueNorth chip from DARPA and IBM
Reference: (Akopyan et al., 2015)
TrueNorth is a neuromorphic computing chip built by DARPA and IBM. To mimic neurons, it uses circuit modules for which inputs must surpass a threshold in order to allow a signal to pass. The chip can emulate any desired configuration of up to a million neurons with over 250 million synapses in real time while consuming far less power than traditional computing devices. Multiple chips can be interconnected to enable larger simulations.

2016
32 – Electrode array restores some tactile sensations
Reference: (Flesher et al., 2016)
Electrode array stimulation of specific parts of the somatosensory cortex partially restored tactile sensations within the hand of an injured patient over a period of six months. Furthermore, the intensity of sensory stimuli was adjustable through modulating the amplitude of the electrode array inputs, suggesting that complicated forms of sensory information (i.e. pressure differences) might be restored using this strategy. Results from this study might be used to engineer neuroprotheses which give tactile feedback.

33 – Expansion FISH
Reference: (F. Chen et al., 2016)
Fluorescent in situ hybridization (FISH) was combined with expansion microscopy to facilitate imaging of spatial localization of RNAs within neural tissue. By using a chemical linker to attach intracellular RNAs to the swellable polymer network, the RNAs maintained their spatial locations. FISH staining was then employed after expansion.

34 – Founding of ReadCoor
Reference: (Wyss Institute, 2016)
Fluorescent in situ sequencing (FISSEQ), a method which allows spatially-resolved sequencing of RNAs within intact tissue samples, was invented at Harvard’s Wyss Institute (see highlight #18). After further development, this technology led to a startup called ReadCoor which seeks to use FISSEQ tools for biomedical and connectomics applications. ReadCoor is employing FISSEQ to find the spatial locations and genetic identities of pathogens within infected tissue as a strategy to combat child mortality in developing countries. Another application that ReadCoor pursues is FISSEQ-based evaluation of how drug candidates may operate in the context of spatial tissue gene expression in order help discover better drugs (e.g. nucleic acid therapeutics for treating rare brain diseases). Finally, ReadCoor is working with a governmental research program called the IARPA MICrONS project towards applying FISSEQ to connectomics as a way of informing the design of biomimetic artificial intelligence.

35 – Neural dust
Reference: (Seo et al., 2016)
Small (~1 mm) implantable biosensors called neural dust were made to allow wireless neural recording and were tested successfully in rats. Neural dust motes consist of two recording electrodes, a transistor, and a piezoelectric crystal. They receive external power from ultrasound and allow recording of neuronal activity via disruptions to the piezoelectric crystal’s reflection of the ultrasound waves. In the future, neural dust could be miniaturized further using custom electronic components.

36 – The China Brain Project
Reference: (Poo et al., 2016)
Launched to help understand cognition, complete mesoscopic brain maps, develop new tools for neuroscience, and develop treatments and diagnostics for brain disease, the China Brain Project will take place from 2016 to 2030. The project uses macaques as a central model animal since China has existing infrastructure which takes advantage of the natural macaque population.

2017
37 – Automated patch-clamp robot for in vivo neural recording
Reference: (Suk et al., 2017)
A robotic system for automating patch-clamp recordings of individual neurons was made and tested in vivo using mice. The robot continuously images its target via two-photon microscopy and uses an imagepatching algorithm to control the patch-clamp pipette and compensate for movement in the tissue. The robot achieves similar data yield to that of skilled human experimenters.

38 – Brain imaging factory in China
Reference: (Cyranoski, 2017)
The HUST-Suzhou Institute for Brainsmatics is an industrial-scale neuroscience facility with 50 machines that automatically slice, stain, and image mouse brains. This allows rapid reconstruction of whole brains at the resolution of light microscopy. Although this resolution is not sufficient to densely reconstruct every neuron, sparse staining of cells enables tracing of long-range projection patterns. The institute has already demonstrated promise by mapping the morphology of a previously unknown neuron which seems to wrap around the entire mouse brain.

39 – Bryan Johnson launches Kernel
Reference: (Regalado, 2017)
Entrepreneur Bryan Johnson invested $100 million to start a neurotechnology company called Kernel which seeks to develop implants that allow recording and stimulation of many neurons at once. Kernel’s short-term goal is to develop treatments for brain disease and its long-term goal is to enhance human intelligence.

40 – DARPA funds research to develop superior BCIs
Reference: (Hatmaker, 2017)
Six research groups received portions of a total award of $65 million from DARPA to develop advanced brain-computer interfaces (BCIs). The research groups included five academic laboratories and a small company called Paradromics Inc. DARPA hopes that this investment may lead to a nickel-sized bidirectional BCI which can interact with up to a million individual neurons.

41 – Early testing of hippocampal prosthesis in humans
Reference: (Song, She, Hampson, Deadwyler, & Berger, 2017)
The MIMO algorithm (see hippocampal prosthesis from highlight #8) was used on two human epilepsy patients with implanted recording and stimulation electrodes. The setup recorded from parts of the hippocampus, processed the spiking data, and stimulated other parts of the hippocampus. When assisted by this artificial system, the patients performed significantly better on memory tasks than when the system was turned off, demonstrating that the prosthesis can encode and restore memory.

42 – Elon Musk’s NeuraLink startup
Reference: (Etherington, 2017)
Elon Musk started a neurotechnology company called NeuraLink, a venture which is working on superior brain-computer interfaces (BCIs). NeuraLink plans to initially develop BCIs for clinical applications, but its ultimate goal is to enhance human cognition in order to keep up with artificial intelligence.

43 – Facebook announces effort to build brain-computer interfaces
Reference: (Constine, 2017)
At a company-run conference, Facebook revealed research towards non-invasive BCIs which may eventually allow users to type 100 words per minute using thought alone. Facebook’s team is also working on a wearable device for “skin hearing” which involves vibrating actuators. Test subjects have so far been able to recognize up to nine words using the devices and Facebook hopes to vastly increase these capabilities.

44 – Genome editing in the mammalian brain
Reference: (Nishiyama, Mikuni, & Yasuda, 2017)
Genome editing in the brain has historically been difficult due to most neurons existing in a postmitotic state, preventing homology-directed repair from facilitating insertion of DNA. However, a new technique in which adeno-associated viruses (AAVs) are used to deliver CRISPR-Cas9 systems may overcome this problem. AAVs delivered a ssDNA sequence encoding a guide RNA to transgenic host animals which expressed Cas9, enabling targeted genome editing. Furthermore, minimal off-target effects occurred (1.9%).

45 – Hardware for accelerated and biologically realistic simulations
Reference: (Schemmel, Kriener, Müller, & Meier, 2017)
Scientists working under the Human Brain Project extended the BrainScaleS neuromorphic hardware to help emulate biologically realistic neurons at a speed 1,000 times faster than biological real-time. The hardware’s design incorporates circuit blocks which correspond to compartments representing segments of dendritic trees. A system of switches controls the organization and conductances of these compartments. The compartments can emulate dendritic sodium spikes, dendritic NMDA spikes, or dendritic calcium spikes. Any electronic neuron in the chip can receive up to 16,000 user-configured inputs. Electronic neurons can also communicate between multiple chips. Furthermore, the chip contains hardware systems for mimicking neurological plasticity.

46 – Human Brain Project analyzes tissue using algebraic topology
Reference: (Reimann et al., 2017)
Investigators under the Human Brain Project analyzed their previously reconstructed ~31,000 neuron cortical microcircuit using algebraic topology methods. First, the microcircuit was converted to a digraph. As compared to controls, many directed cliques (complete directed subgraphs) with large numbers of neurons were found in this digraph. By examining these directed cliques using topological metrics, the scientists derived insights on the flow of information in the network. Experimental patch-clamp data supported these insights and the presence of similar patterns in the C. elegans connectome suggested that the results may generalize across species.

47 – Iterative expansion microscopy
Reference: (Chang et al., 2017)
Iterative expansion microscopy (iExM) was developed to facilitate isotropic enlargement of tissue beyond 4x. The iExM process involves performing standard tissue expansion (see highlight #23), reinfusing the expanded sample with a second gel, crosslinking the tissue to the second gel, dissolving the first gel, and carrying out water-mediated expansion again. Using iExM, expansion factors of 20x can be achieved, allowing effective resolutions of about 25 nm when using diffraction-limited light microscopy. At these resolutions, it is possible to visualize detailed structures of cytoskeletal assemblies and other subcellular protein distributions. Furthermore, iExM may enable nanoscale connectomics using light microscopy.

48 – Neuropixels
Reference: (Jun et al., 2017)
Neuropixels is a probe which allows simultaneous recording from hundreds of neurons with high spatiotemporal resolution. Its recording shank is one centimeter long and has 384 recording channels. The accompanying apparatus is small (6×9 mm base and a data transmission cable), enabling recording in freely moving animals.
![]()
49 – Simulation of rat CA1 region
Reference: (Bezaire, Raikov, Burk, Vyas, & Soltesz, 2016)
Multicompartmental Hodgkin-Huxley models of 338,740 neurons including pyramidal cells and several interneuron types were equipped with approximate connectivity patterns based on data from the biological CA1 region of rat brains (over 5 billion synapses were present). Different cell types received different numbers of synapses at specified distances from the soma. External inputs to the region were also estimated based on biological data. Supercomputers were used to carry out the simulation, but even so, one second of the simulation took about 4 hours to complete. As in the biological CA1 region, certain types of oscillations occurred. Using the simulation, parvalbumin cells and neurogliaform cells were identified as central drivers of these oscillations (this was not previously known).

50 – The $100 billion Softbank Vision Fund
Reference: (Lomas, 2017)
Masoyoshi Son (CEO of Softbank) announced a plan to partner with multiple large companies and raise $100 billion to invest in artificial intelligence. By the end of 2017, the Vision Fund had successfully reached this goal. Son hopes to play a part in bringing the technological singularity to fruition in order to help tackle humanity’s greatest challenges.

51 – UltraTracer enhances existing neuronal tracing software
Reference: (Peng et al., 2017)
UltraTracer is an algorithm which can improve the efficiency of existing neuronal tracing software for handling large datasets (e.g. hundreds of billions of voxels or more). It typically enhances performance by 3-6 times, though some programs can have their speed increased 10-30 times. Ultratracer is opensource and available as a plugin for the Vaa3D software platform.

52 – Whole-brain electron microscopy in larval zebrafish
Reference: (Hildebrand et al., 2017)
A millimeter-long region encompassing the brain and spinal cord of a larval zebrafish was imaged using serial-section electron microscopy (ssEM). Using a modified version of the CATMAID neuronal tracing software, all 2,589 myelinated axons in the dataset were reconstructed along with some of the associated soma and dendrites. Unfortunately, only 834 of the myelinated axons were traced back to cell bodies. These ssEM data were also compared to two-photon functional imaging data from the same specimen to facilitate better understanding of the relationship between structure and function.

2018
53 – Data-informed model of neuron densities and positions in mouse brain
Reference: (Erö, Gewaltig, Keller, & Markram, 2018)
As part of the Blue Brain Project, a virtual 3D model of cell locations in the mouse brain was constructed. Nissl staining data and mouse brain region demarcations from the Allen Institute were used to computationally estimate cell densities across 737 discrete brain structures. Next, an algorithm randomly generated exact cell positions as constrained by the density map and estimated total cell counts. With these constraints, the estimated cell positions across the mouse brain demonstrated a good match when compared to the real biological data, though some minor sources of error did occur. Data from the Allen Institute describing neuronal gene expression were used to label the cells in the algorithmically generated map as excitatory neurons, inhibitory neurons, astrocytes, oligodendrocytes, and microglia. These labels were generated by an algorithm which was informed by the data from the gene expression atlas. The distributions of modulatory neurons (dopaminergic, cholinergic, and serotoninergic) within selected regions were also incorporated into the 3D atlas. Analyzing this virtual 3D atlas model of mouse brain cells can yield insights into the cellular organization of the mouse brain in a way that would not be possible with only the raw datasets. The approach to constructing this model allows for further datasets to be easily integrated into its framework. Furthermore, the 3D atlas model is freely available online and comes with a user-friendly interface. Finally, this model is planned to be used as a spatial foundation for constructing biophysically detailed whole-brain simulations.

54 – Detailed simulation of visual cortex layer IV
Reference: (Arkhipov et al., 2018)
Using 10,000 multicompartmental Hodgkin-Huxley neurons (100-200 compartments each) and 35,000 leaky-integrate-and-fire neurons, a simulation of layer IV of mouse primary visual cortex was built. The multicompartmental neuron models included three distinct excitatory cell types and two distinct inhibitory cell types. In addition, a greatly simplified model of lateral geniculate nucleus (LGN) processing was made to allow visual sensory input to enter the system. This LGN model consisted of linear-nonlinear Poisson filters which were pooled to create orientation-selective inputs at the appropriate cortical cells. Connectivity was first determined as random except for higher likelihoods of connections forming between closer cells. Next, connectivity was modified by assigning each excitatory cell a preferred stimulus orientation and giving higher synaptic weights to connections between cells with more similar stimulus preferences. Numbers of synapses were based on values from the literature and confirmed via electron microscopy. Externally-generated waves of background activity were implemented in the model to reflect inputs from other brain areas. Plasticity was neglected since the simulation only covered a few seconds of real time. The system was exposed to simulated visual stimuli such as gratings and natural movies. Though some deficiencies occurred (e.g. no direction selectivity and too rapid of responses to large flashes of light), the simulation reproduced many features of biological neural activity such as orientation selectivity, magnitudes of responses to gratings, distributions of firing rates, and others. Furthermore, changing various properties of the system led to insights on what kinds of details are important for brain simulations.

55 – EEG-based facial image reconstruction
Reference: (Nemrodov, Niemeier, Patel, & Nestor, 2018)
By collecting EEG data associated with participants viewing images of faces, neural correlates of facial processing were determined. Using a mathematical algorithm, these data were used to reconstruct images of the faces and compare them to the original stimulus images. The most accurate EEG-based reconstructions were based on data from multiple people, but single person data still yielded statistically significant accuracy. Beyond its neurotechnological applications, this process gave insights on the cognitive steps in facial perception.

56 – Further clinical testing of hippocampal prosthesis in humans
Reference: (Hampson et al., 2018)
In more clinical testing of the hippocampal prosthesis (see highlight #41 for early clinical testing), twenty-one epilepsy patients took part in the experiments. Seventeen patients underwent hippocampal recording to facilitate optimization of the MIMO model and eight patients received hippocampal stimulation to improve their memory abilities. To accomplish this, electrodes were implanted for recording from the CA3 region of the hippocampus and stimulation at the CA1 region of the hippocampus. The hippocampal prosthesis system granted an average improvement of 35% on behavioral memory tasks.

57 – Human telepathy using BrainNet
Reference: (Jiang et al., 2018)
A form of human telepathic communication was carried out by taking EEG recordings from two people (called senders) while they played a Tetris-like game and transmitting signals derived from the recordings to a receiver person’s occipital cortex via transcranial magnetic stimulation (TMS). With this setup, the third individual was able to make decisions in the game without seeing the screen. To convey information, the senders were told to focus on either a higher or a lower intensity light. These lights were positioned on either side of the computer screens. When the receiver’s occipital cortex underwent TMS stimulation, the receiver perceived a mental image of a flash of light. The receiver could distinguish between the different intensities corresponding to the lights as seen by the senders. Using only this telepathic system, the receiver made the correct gameplay decisions 81% of the time. Five groups of three people took part in this study.

58 – Improved spatial RNA sequencing using STARmap
Reference: (Wang et al., 2018)
Spatially-resolved transcript amplicon readout mapping (STARmap) was developed to sequence RNA within brain tissue and so acquire 3D spatial information regarding RNA localization. This method is related to FISSEQ (see highlights #18 and #34), but it exhibits higher throughput and can be applied to thicker tissue volumes. To carry out STARmap, fixed tissue is first infused with a hydrogel matrix which helps enhance the optical translucency of the sample. DNA amplicons are covalently linked to this matrix. With the help of reverse transcriptase, these amplicons carry out replication of local mRNAs to amplify the signal-to-noise ratio in subsequent steps. Each amplicon contains a gene-unique barcode sequence. Next, the modified tissue sample is washed with fluorescently labeled DNA probes of distinct colors and imaged after each wash. By observing which spots fluoresce over the course of six or seven cycles, the sequences of the barcodes associated with RNAs at spatially defined locations are found. Finally, the RNAs are identified via these barcode sequences. STARmap has been used to quantify the spatial expression of up to 1020 genes across 30,000 cells in a volume of mouse cortex.

59 – Map of all neuronal cell bodies within mouse brain
Reference: (Murakami et al., 2018)
To help create an atlas of cell bodies in the mouse brain, a new chemical tissue treatment method called CUBIC-X was developed via intensive screening of various reagents. CUBIC-X optically clears mouse brains as well as physically enlarging them by a factor of 2x. Using CUBIC-X and light-sheet fluorescence microscopy, whole mouse brains were imaged at subcellular resolution. Novel software was written to identify all individual cellular nuclei. With these tools, an atlas of the spatial locations of all cell bodies within several adult and developing mouse brains was constructed. This atlas was made publicly available online.

60 – Nanowire device as an artificial synapse
Reference: (Milano et al., 2018)
To aid future neuromorphic computing systems, a nanowire-based device was synthesized to act as a memristive synapse. The device consists of a ZnO nanowire bridging between silver and platinum electrodes on an insulating substrate. When a positive voltage is applied to the silver electrode, Ag+ ions migrate along the nanowire and undergo deposition, creating a conductive bridge. Each time voltage is applied, conductance of this bridge increases as more silver is added to the wire. In this way, the system mimics synaptic potentiation. Pulses with negative voltage can reset the nanowire bridge’s conductance back to its initial state. Short-duration pulses destabilize the silver on the nanowire and cause the conductance to decay. By using certain amplitudes and frequencies of voltage pulses, the kinetics of bridge formation and decay can mimic the kinetics of synapses.

61 – Precise optogenetic manipulation of many neurons simultaneously
Reference: (Mardinly et al., 2018)
By constructing a holographic optical system called 3D-SHOT and engineering superior optogenetic channels, it was possible to gain precise spatiotemporal control over complex patterns of neural activity in living mice and with in vitro neuronal setups. For the case of living mice, 21 random neurons were targeted with spots of light inside of a brain volume of 250×250×200 μm3 at a depth of 113-323 μm below the pial surface. To generate these spots of light, 3D-SHOT uses a spatial light modulator to distribute a laser beam (with two-photon excitation) onto multiple targets. The new optogenetic channels include an excitatory channel called ST-ChroME and an inhibitory channel called IRES-ST-eGtACR1. These channels generate strong photocurrents over short timescales and are localized to the somas of host neurons, facilitating better optogenetic precision.

62 – Software for making large-scale brain simulations more efficient
Reference: (Jordan et al., 2018)
Large-scale simulations of networks of spiking neurons require enormous computational resources due to the challenge of working with vast numbers of synaptic connections. To ameliorate this difficulty, a novel software architecture which more efficiently routes information in large networks of neurons (i.e. more than 109 neurons) was designed. The software operates by first storing connection information in postsynaptic neurons and subsequently (just prior to the start of a simulation) constructing the presynaptic infrastructure based on the postsynaptic connection information. For handling neural spike communication, the software retrieves target information from the presynaptic infrastructure and uses this information to modify a four-dimensional array called the spike buffer. The spike buffer communicates spikes by instructing a message passing interface (MPI). An MPI is a way of transporting information between different parts of a high-performance computing system. By using the spike buffer to inform the MPIs, only the MPI processing that is strictly necessary will take place, making it so that less computational resources are required. This software may help large-scale brain simulations run on exascale computers. Furthermore, the software can also run on much smaller computers without penalizing efficiency. Although the software is highly scalable for single-compartment spiking neuron models, it is not optimized for biologically realistic multicompartmental simulations or models of neuronal plasticity, so more work will need to be carried out to incorporate these biological constraints.

63 – SpiNNaker neuromorphic supercomputer
Reference: (Brown, Chad, Kamarudin, Dugan, & Furber, 2018)
Under funding from the Human Brain Project, a neuromorphic supercomputer called SpiNNaker was constructed to simulate spiking neuronal networks in real time. Its neuromorphic architecture makes SpiNNaker uniquely optimized for carrying out these kinds of tasks. It consists of a mesh of nodes, each with eighteen ARM9 processors. Small information packets which carry the equivalent of a neuronal spike are rapidly transmitted among the nodes via a hardware-based routing system. SpiNNaker can simulate up to one billion single-compartment neuron models (e.g. integrate-and-fire neurons, Izhikevich neurons, etc.) It can also simulate multicompartmental neurons, but each neuron then requires more of SpiNNaker’s resources.

64 – Transcriptomic cell type classification across mouse neocortex
Reference: (Tasic et al., 2018)
Using single-cell RNA sequencing, gene expression across 23,822 visual cortex and motor cortex cells was characterized. Dimensionality reduction and clustering methods enabled classification of these cells into 133 transcriptomic cell types. By injecting adeno associated viruses which carried genes for fluorescent proteins, neuronal projections for a subset of the sequenced cells underwent tracing. This established correspondences between projection patterns and transcriptomic identities.

65 – Upconversion nanoparticles for noninvasive optogenetic stimulation
Reference: (S. Chen et al., 2018)
Upconversion nanoparticles absorb two or more photons of lower energy and then emit a single photon of higher energy. To achieve noninvasive optogenetic stimulation, upconversion nanoparticles were injected into the brains of living mice. Externally applying near-infrared light (which is low energy and penetrates tissue) was converted into blue light inside the mouse brains by the nanoparticles, causing activation of the excitatory optogenetic ChR2 channel. Another type of upconversion nanoparticle was similarly used to generate green light within living mouse brains, causing activation of the inhibitory optogenetic Arch channel. As a proof-of-concept, this technique was used to trigger hippocampally-encoded fear memories in the mice.

66 – Whole-brain Drosophila image data via electron microscopy
Reference: (Zheng et al., 2018)
An entire Drosophila brain volume was imaged using serial-section transmission electron microscopy at a voxel size of 4×4×40 nm, providing the data needed to reconstruct a complete map of the fly’s brain. These data take up 106 TB of storage space. Though the process of reconstructing a Drosophila connectome from this dataset has not yet been completed, substantial progress has been made.

67 – X10 expansion microscopy
Reference: (Truckenbrodt et al., 2018)
X10 microscopy was developed to enable isotropic enlargement of tissue beyond 4x without necessitating the challenging process of iterative expansion microscopy (see highlight #47). Using some alternative chemical reagents compared to traditional expansion microscopy, a 10x expansion factor is achieved and resolutions of around 25 nm are possible with many types of conventional light microscopes. Similar to iterative expansion microscopy, X10 microscopy allows imaging of molecular assemblies as well as fine neuronal structures like synapses.

2019
68 – Algebraic topology for morphological classification of pyramidal cells
Reference: (Kanari et al., 2019)
Scientists working under the Human Brain Project employed a topological data analysis method to objectively classify pyramidal cells based on their morphologies. Cell type classification is often performed by experts making subjective evaluations of neuronal morphology, so topological data analysis is an overall more consistent and rigorous technique. An algorithm called the topological morphology descriptor (TMD) facilitated conversion of 3D reconstructions of pyramidal cells from distinct cortical layers into barcodes describing the branching patterns of their dendritic trees. Topological distances between the barcodes were computed using several mathematical metrics defined by previous literature. Next, a support vector machine (a type of machine learning algorithm) was used to cluster neurons into classes based on the computed distances between barcodes. With this process, 17 distinct types of pyramidal cells in rat somatosensory cortex were identified in an automated and unbiased fashion.

69 – Approximate model of mouse neocortical connectome
Reference: (Reimann et al., 2019)
By using connectivity rules derived from a variety of datasets, scientists working under the Blue Brain Project built a computationally-generated approximate model of the mouse neocortical connectome. To start, about 10 million morphological reconstructions of neurons (many copies of 55 distinct morphologies) were placed into a virtual 3D space with the shape of the mouse cortex. A data-derived atlas of neuron densities by brain region (see highlight #53) was employed to determine the densities of excitatory and inhibitory neurons throughout the volume. Local connectivity patterns were decided using an algorithm which took into account layer-specific densities of each neuron type, the lengths of axons for each neuron type, the densities of synaptic boutons coming from the axons of each neuron type, the average number of synapses per connection between neurons for each pair of morphological neuron types, and the connection probability for each pair of morphological neuron types (as in highlight #24). Long-range connectivity patterns and projection strengths were decided using a mesoscale connectivity dataset (from viral tracing experiments). This dataset included connection strengths between source and target regions of the mouse brain as well as layer-specific connection profiles. Further analysis of the mesoscale connectivity dataset’s structure also allowed incorporation of variations in the finer spatial preferences of projections within each source and target region. The MouseLight dataset (see highlight #85) was used to estimate what types of neurons were present in each projection and to set rules on the number and identity of distinct brain regions targeted by individual neurons from a given source region. These analyses were used to constrain the long-range connections of reconstructed neurons across the cortical model. As with the local connectivity patterns, subcellular target locations of the long-range projections were defined according to a data-supported algorithm. The resulting model of the mouse neocortical connectome will later be used to support large-scale biophysical simulations of the mouse cortex.

70 – Automated electron microscopy of a 1 mm3 volume of mouse cortex
Reference: (Yin et al., 2019)
Allen Institute scientists developed an automated serial-section transmission electron microscopy (ssTEM) pipeline which features parallel operation of five TEMs at once. Using this system, a 1 mm3 region of mouse cortex underwent high-resolution imaging (4×4×40 nm voxels) over a period of less than six months. An automated tape collecting ultramicrotome (ATUM) was employed to make the 40 nm slices of tissue (see highlight #8) and GridTape (see highlight #76) was utilized to collect the sections for subsequent imaging. Numerous customized hardware and software components were used to facilitate continuous automated operation through the day and night and to ensure that quality data could be efficiently acquired.

71 – Decoding sentences from brain activity
Reference: (Anumanchipalli, Chartier, & Chang, 2019)
Electrocorticography and machine learning were utilized to translate brain activity into the correct sentences. By training neural networks to compare audio data with facial movements during speech and then to compare facial movements during speech with brain activity, a machine learning tool was developed to synthesize audio based on brain activity. With this tool, patients were able to silently mime sentences and have the system speak for them. Though some errors occurred in the translation from brain activity to speech, the system could still be very helpful to people with vocal disorders.

72 – Detailed reconstruction of a larger volume of mouse cortex
Reference: (Motta et al., 2019)
After using serial block-face electron microscopy (SBEM) to image a volume of layer IV mouse somatosensory cortex with dimensions of ~62×95×93 μm, a combination of automated image processing and human annotation allowed full segmentation of the volume and reconstruction of its connectome. As of the publication of this study, this volume is the largest tissue region for which a connectome has been fully reconstructed. Automated segmentation, synapse detection, and other analysis were first carried out via a suite of electron microscopy image processing software. Artificial intelligence algorithms were developed to direct human annotators to regions of likely error within the automatically processed dataset. An optimized user-interface was designed to allow the annotators to rapidly fix these errors. This analysis took about 4,000 human work hours as distributed among 87 people, a rate 20 times faster than had been achieved in previous studies. The connectome of the volume included 6,979 presynaptic neurites and 3,719 postsynaptic neurites with a total of 153,171 synapses. After fully segmenting the volume and reconstructing its connectome, numerous further analyses were carried out to reveal quantitative insights on the organization of the dataset’s neuronal structures.

73 – Expansion lattice light-sheet microscopy
Reference: (Gao et al., 2019)
By combining lattice light-sheet microscopy with expansion microscopy, the entire Drosophila brain as well as subvolumes of mouse cortex underwent imaging at 60×60×90 nm resolution (though target structures were sparsely labeled). For the 340×660×90 μm Drosophila brain, all 110 dopaminergic neurons were immunostained with one color and all ~40 million presynaptic boutons were immunostained with a second color. Eight of the dopaminergic neurons were manually traced. For the mouse cortex subvolumes (100×150×150 μm and 320×280×60 μm), the morphological parameters of fluorescently labeled organelles as well as the structural properties of fluorescently labeled myelin sheaths were quantified. Since the lattice light-sheet microscope’s field of view was only 160 μm, hundreds or even thousands of tiles were computationally stitched together using customized software to reconstruct the full images.

74 – Fruit Fly Brain Observatory
Reference: (Ukani et al., 2019)
The Fruit Fly Brain Observatory (FFBO) is an opensource software platform for integrating Drosophila-related databases and computational modeling. The FFBO hosts components including NeuroArch, Neurokernel, NeuroNLP, and NeuroGFX. NeuroArch contains all of the connectomic and anatomical data from existing databases such as FlyCircuit and Janelia’s electron microscopy reconstructions as well as some electrophysiological data from various parts of the fly brain. Neurokernel is an engine which uses GPUs to facilitate interconnection of smaller models of distinct parts of the fruit fly brain into larger unified simulations. NeuroNLP provides an online graphical user interface which allows navigation of Drosophila neural circuit data (e.g. connectomic, cell-type, electrophysiology, etc.) and includes a natural language query system. NeuroGFX features tools for bringing together circuit diagrams, biological data, computational models, and visualization of simulation results. Together, these components comprise a software ecosystem that facilitates integrative computational studies of the Drosophila brain.

75 – Gas cluster ion beam scanning electron microscopy
Reference: (Hayworth et al., 2019)
To help make feasible the imaging of millimeter-scale volumes of brain tissue at 10 nm3 voxel size or better, a new form of scanning electron microscopy called gas cluster ion beam scanning electron microscopy (GCIB-SEM) was developed. GCIB-SEM starts with ultramicrotome-based cutting of relatively thick (0.5-1.0 μm) sections of tissue which are mounted on a silicon wafer. This step can be automated via ATUM-SEM (see highlight #8). Next, the sections are irradiated by a flood of electrons to increase their conductivity, mitigating charge accumulation during imaging. SEM imaging is carried out on these sections and then a beam of energetic argon atom clusters mills away a thin layer of tissue (about 10 nm). The imaging and milling steps are repeated until the thick sections are gone. By comparison to past ion beam milling methods, the gas cluster ion beam has properties which make it possible to image larger areas more rapidly. The milling step of GCIB-SEM is fast enough to mill 1 mm3 of tissue per month, making the technique highly efficient compared to past methods. GCIB-SEM was also demonstrated to be compatible with multibeam SEM (see highlights #26 and #27), giving it the potential for very high throughput. As a proof-of-concept, GCIB-SEM was tested on several small tissue volumes. Computational methods facilitated 3D reconstruction and correction of minor discrepancies between the sections. The high resolution of GCIB-SEM should make it possible to achieve fully automated computational segmentation of imaged tissue.

76 – GridTape for automated transmission electron microscopy
Reference: (Graham et al., 2019)
GridTape is a substrate for carrying tissue sections during transmission electron microscopy (TEM). It enables using an automated tape collecting ultramicrotome (ATUM) with TEM. ATUM was previously compatible only with scanning electron microscopy (see highlight #8). This is important since TEM can achieve higher spatial resolution and much higher signal-to-noise ratios compared to SEM. To make GridTape, lasers were used to mill regularly spaced holes into aluminum-coated polyimide tape. The tape was then coated with a very thin electron-lucent film which spanned the holes. An automated TEM sample stage for GridTape was also built, allowing rapid positioning of tissue sections beneath the electron beam and shifting of sections to capture montages of the regions of interest.

77 – Injectable nanoparticles give mice infrared vision
Reference: (Ma et al., 2019)
Subretinal injections of upconversion nanoparticles gave mice the ability to perceive complex shapes in the near-infrared spectrum. These nanoparticles were linked to a protein which binds sugars on photoreceptor cell surfaces. Since upconversion nanoparticles absorb longer wavelengths of light and emit shorter wavelengths of light, this allowed the mice to perceive near-infrared light as if it was visible light. Furthermore, treated mice retained their ability to see visible light while gaining the ability to see near-infrared light. Minor side effects occurred (cataracts and corneal opacity) but disappeared after two weeks. The mice could still distinguish complex near-infrared shapes ten weeks after treatment.

78 – Morpho-electric cell type classification in mouse visual cortex
Reference: (Gouwens et al., 2019)
By collecting electrophysiological data from 1,938 neurons and making morphological reconstructions of 461 neurons (with an overlap of 452 neurons) and carrying out unsupervised clustering methods on the data, neurons were classified into distinct cell types. There were 17 electrophysiological cell types, 38 morphological cell types, and 46 morpho-electric cell types. The morpho-electric cell types were derived from joint clustering on the combined electrophysiology and morphology datasets. Electrophysiology data came from patch-clamp recordings of a diverse array of cells labeled with fluorescent proteins. To acquire the morphology data, the recording pipettes were used to fill targeted neurons with biocytin so as to facilitate sparse fluorescence-based reconstruction. Morpho-electric cell type classification may aid brain simulations since it could facilitate prediction of electrophysiological properties from morphological data.

79 – NeuraLink reveals brain-machine interface prototype
Reference: (Musk, 2019)
Elon Musk’s NeuraLink company developed a prototype brain-machine interface system which consists of flexible threads carrying recording electrodes, a small electronic device for collecting and processing the recording data from the threads, and a surgical robot that inserts these threads into brain tissue. The threads are at most 6 μm in thickness, are made from polyimide-coated gold, each have 32 gold electrodes with chemical surface modifications to improve their conductivity, and connect to an application specific integrated circuit (ASIC). A small electronic device with a USB port contains several of these ASICs as well as some other electronic components. The device rejects noise, amplifies neural signals, and digitizes the signals before the data are transmitted via the USB port. In total, the device can use its threads to record from up to 3,072 neurons at once. The device only consumes ~6 mW of power. To implant the threads, a neurosurgical robot was designed. The robot includes stereoscopic cameras and light sources to track the 3D position of the threads as they undergo insertion into the desired anatomical region of the brain. This also allows the surgical robot to avoid damaging microvasculature as it inserts the threads. Though the robot can operate automatically, surgeons retain control and can make adjustments if needed. The surgical robot and brain-machine interface system have been tested on rats.

80 – Neural probes that resemble real neurons
Reference: (Yang et al., 2019)
Neural recording devices shaped similarly to biological neurons were constructed. The probes (called NeuE probes) consist of soma-like platinum recording electrodes and neurite-like filaments made from a gold wire coated in a polymer insulator. The sizes of these components are similar to the sizes of real somas and neurites. In addition, the filaments exhibit similar flexibility compared to biological axons. When tested in living mouse brains, the NeuE probes neither depleted surrounding neurons nor induced microglial responses, indicating high biocompatibility. NeuE probes with 16 electrodes were implanted and demonstrated stable recordings over periods of 90 days.

81 – Optogenetic inception of memories to guide vocal learning in songbirds
Reference: (Zhao, Garcia-Oscos, Dinh, & Roberts, 2019)
Using an adeno-associated viral vector, an axon-targeted ChR2 ion channel (see highlight #1) was expressed at the interfacialis of the nidopallium region of the songbird brain. With this optogenetic setup, light pulses mimicking some of the rhythms of birdsong allowed vocal learning in young songbirds who were isolated from adult songbird tutors. Different patterns of light pulses resulted in distinct song behaviors as the young songbirds matured. Because of these pattern-dependent variations in how the songs of the young birds developed over time, the light pulses may have created memories which guided the songs rather than simply training a specific sequence of motor actions. Optogenetic stimulation was found to override the effects of tutoring from older birds when young birds were exposed to both tutoring and light pulses. In addition, these investigations uncovered insights regarding circuit function in songbird vocal learning.

82 – Partial segmentation of whole-brain Drosophila electron microscopy dataset
Reference: (Li et al., 2019)
Though the entire Drosophila brain was imaged using electron microscopy (see highlight #66), the data still require segmentation to define each neuron as a distinct object and to aid the construction of a connectome. This task has not yet been completed in its entirety. However, an automated algorithm involving flood-filling networks was used to carry out a draft segmentation of the dataset. With this draft, a large number of split errors are present. Nonetheless, these errors can be manually corrected in a tenth of the time it would take to manually trace the neurons from the raw data. It is also expected that new algorithms with superior performance will be developed.

83 – Perovskite nickelates facilitate superior sensing of molecules
Reference: (Zhang et al., 2019)
Using a perovskite nickelate lattice with covalently linked horseradish peroxidase (HRP) enzyme molecules, dopamine release from synapses in a mouse brain slice was measured with high selectivity and sensitivity. Upon stimulation of the dopaminergic neurons in the slice, the HRP reacted with the dopamine and initiated a process of hydrogen transfer onto the perovskite nickelate lattice. Even in a complex medium of artificial cerebrospinal fluid, the device selectively detected dopamine. The device’s sensitivity was high enough that dopamine could be detected at concentrations of 5•10-17 M. A similar device was built for sensing glucose molecules by linking a glucose oxidase enzyme to the perovskite nickelate lattice (rather than HRP). This device was selective enough to respond only to glucose and not to mannose or galactose. It could detect glucose at concentrations of 5•10-16 M.

84 – Pipeline for mesoscale mapping of the marmoset brain
Reference: (Lin et al., 2019)
As part of Japan’s Brain/MINDS project (see highlight #26), an experimental pipeline for mesoscale mapping of the marmoset brain was built. Mesoscale refers to connectivity between anatomical regions. To start, MRI-guided injection of neural tracers was carried out. Each marmoset brain was then sectioned into ~1,700 slices with 20 μm thicknesses. In addition, a set of chemical stains was applied to enhance visualization of neuronal features. Imaging the slices facilitated making 3D reconstructions of the marmoset brains. The MRI data helped correct distortions in these reconstructions and enabled registration into a standard 3D marmoset brain atlas. The fluorescent tracers facilitated making weighted connectivity maps between distinct anatomical regions. With this pipeline, a mesoscale connectivity matrix for the entire marmoset brain might be achieved by 2024.

85 – Reconstruction of 1,000 long-range projection neurons in mouse brain
Reference: (Winnubst et al., 2019)
To reconstruct over 1,000 long-range neuronal morphologies across mouse brains, the animals were first injected with viral vectors carrying genes which enabled sparse fluorescent labeling of neurons. Whole brains were harvested, optically cleared, treated via immunohistochemical methods to amplify the fluorescent signals, and imaged using an automated two-photon microscope and vibratome. The two-photon microscope performed 3D imaging at depths of 250 μm and the vibratome sliced the tissue at depths of 175 μm, giving 75 μm of overlap between layers (to facilitate image stitching). In this way, each whole mouse brain was imaged at 0.3×0.3×1.0 μm voxel size over the course of a week. Each mouse brain supported reconstruction of about 100 neurons. Semi-automated segmentation of the neuronal morphologies was carried out by first performing automated segmentation and then having human annotators correct the software’s errors. With the dataset resulting from this study, the authors gleaned numerous insights into patterns of long-range neuronal projections. The reconstructed neurons can be visualized using a freely available online interface.

86 – Superior connectomes for both C. elegans sexes
Reference: (Cook et al., 2019)
The full connectomes of both the male and the hermaphrodite C. elegans sexes were reconstructed. Though a C. elegans connectome was first completed in 1986, the process was at that time only performed for the C. elegan’s hermaphrodite and not for the male. The early connectome also contains more errors due to the time’s less mature technology and does not describe the weights of synaptic and gap junction links between neurons. In the more recent reconstruction of the male and hermaphrodite C. elegans connectomes, a superior level of detail was achieved. New electron microscopy data were collected when necessary and the 1986 data were digitized and reanalyzed when appropriate. It should be noted that some structures were inferred by combining electron microscopy images from multiple worms. By using a software program called Elegance to facilitate manual annotation of features with single mouse clicks, skeletonized tracings of all of the male and hermaphrodite neurons were made. Elegance allowed annotation of synapses and gap junctions along with their weights (based on the sizes of the synaptic and gap junction structures). The modern investigation also uncovered the neural innervations of C. elegans musculature and other output tissues in the digestive, reproductive, and integumentary systems.

87 – X-ray holographic nano-tomography of neuronal tissue
Reference: (Pacureanu et al., 2019)
A synchrotron radiation source was used to carry out a novel phase contrast x-ray imaging process which achieved 30-100 nm3 voxel sizes on relatively large tissue samples including whole Drosophila brains, a Drosophila leg, and regions of mouse cortex. These high-resolution results were obtained by performing four tomographic acquisitions at different distances from the detector and using a phase retrieval algorithm to make higher-quality reconstructions. Imaging was performed under cryogenic conditions to prevent tissue damage due to the bright x-ray beam. Each x-ray holographic nanotomography scan took about four hours to complete, though some samples required multiple scans and stitching of the resulting images (e.g. the Drosophila leg needed 12 scans). In the Drosophila leg sample, 108 axons were manually skeletonized as well as classified based on morphology. Next, a convolutional neural network was used to perform automated segmentation of the neurons in part of the Drosophila leg. The automated segmentation process took place at a rate of 0.1 s/μm3, about 100 times faster than automated segmentation of electron microscopy data.

88 – X-ray microtomography of whole zebrafish
Reference: (Ding et al., 2019)
Synchrotron-based x-ray microtomography was performed on whole larval and juvenile zebrafish with 0.743 μm3 and 1.43 μm3 voxel sizes respectively. To introduce x-ray contrast, the zebrafish were stained with phosphotungstic acid. The field of view used with the larval zebrafish was 1.5 mm3 (10x objective lens) and the field of view used with the juvenile zebrafish was 3.0 mm3 (5x objective lens). Monochromatic acquisitions took 20 minutes and polychromatic acquisitions took 20 seconds, though the polychromatic imaging exhibited somewhat poorer signal-to-noise ratios. Each zebrafish was imaged over the course of 3-5 acquisitions depending on its size. The numbers and distributions of neuronal cell nuclei in each zebrafish brain were detected and quantified using an automated supervised learning algorithm. This allowed visualization with a heat map to show relative densities of neurons within different zebrafish brain regions. Several other anatomical features, including some anatomical features in zebrafish with pathology-causing mutations, were quantified using the tomographic imaging data. This x-ray microtomography method enables high-throughput phenotyping of small organisms with submicron detail.

References
Ahrens, M. B., Orger, M. B., Robson, D. N., Li, J. M., & Keller, P. J. (2013). Whole-brain functional imaging at cellular resolution using light-sheet microscopy. Nature Methods, 10, 413. Retrieved from https://doi.org/10.1038/nmeth.2434
Akopyan, F., Sawada, J., Cassidy, A., Alvarez-Icaza, R., Arthur, J., Merolla, P., … Modha, D. S. (2015). TrueNorth: Design and Tool Flow of a 65 mW 1 Million Neuron Programmable Neurosynaptic Chip. IEEE Transactions on Computer-Aided Design of Integrated Circuits and Systems, 34(10), 1537–1557. https://doi.org/10.1109/TCAD.2015.2474396
Anumanchipalli, G. K., Chartier, J., & Chang, E. F. (2019). Speech synthesis from neural decoding of spoken sentences. Nature, 568(7753), 493–498. https://doi.org/10.1038/s41586-019-1119-1
Arkhipov, A., Gouwens, N. W., Billeh, Y. N., Gratiy, S., Iyer, R., Wei, Z., … Koch, C. (2018). Visual physiology of the layer 4 cortical circuit in silico. PLOS Computational Biology, 14(11), e1006535. Retrieved from https://doi.org/10.1371/journal.pcbi.1006535
Berger, T. W., Song, D., Chan, R. H. M., Marmarelis, V. Z., LaCoss, J., Wills, J., … Granacki, J. J. (2012). A Hippocampal Cognitive Prosthesis: Multi-Input, Multi-Output Nonlinear Modeling and VLSI Implementation. IEEE Transactions on Neural Systems and Rehabilitation Engineering, 20(2), 198–211. https://doi.org/10.1109/TNSRE.2012.2189133
Berning, S., Willig, K. I., Steffens, H., Dibaj, P., & Hell, S. W. (2012). Nanoscopy in a Living Mouse Brain. Science, 335(6068), 551 LP-551. Retrieved from http://science.sciencemag.org/content/335/6068/551.abstract
Bezaire, M. J., Raikov, I., Burk, K., Vyas, D., & Soltesz, I. (2016). Interneuronal mechanisms of hippocampal theta oscillations in a full-scale model of the rodent CA1 circuit. ELife, 5, e18566. https://doi.org/10.7554/eLife.18566
Boyden, E. S., Zhang, F., Bamberg, E., Nagel, G., & Deisseroth, K. (2005). Millisecond-timescale, genetically targeted optical control of neural activity. Nature Neuroscience, 8, 1263. Retrieved from http://dx.doi.org/10.1038/nn1525
Brown, A. D., Chad, J. E., Kamarudin, R., Dugan, K. J., & Furber, S. B. (2018). SpiNNaker: Event-Based Simulation—Quantitative Behavior. IEEE Transactions on Multi-Scale Computing Systems, 4(3), 450–462. https://doi.org/10.1109/TMSCS.2017.2748122
Chang, J.-B., Chen, F., Yoon, Y.-G., Jung, E. E., Babcock, H., Kang, J. S., … Boyden, E. S. (2017). Iterative expansion microscopy. Nature Methods, 14(6), 593–599. https://doi.org/10.1038/nmeth.4261
Chen, F., Tillberg, P. W., & Boyden, E. S. (2015). Expansion microscopy. Science, 347(6221), 543 LP-548. https://doi.org/10.1126/science.1260088
Chen, F., Wassie, A. T., Cote, A. J., Sinha, A., Alon, S., Asano, S., … Boyden, E. S. (2016). Nanoscale imaging of RNA with expansion microscopy. Nature Methods, 13, 679. Retrieved from http://dx.doi.org/10.1038/nmeth.3899
Chen, S., Weitemier, A. Z., Zeng, X., He, L., Wang, X., Tao, Y., … McHugh, T. J. (2018). Near-infrared deep brain stimulation via upconversion nanoparticle–mediated optogenetics. Science, 359(6376), 679 LP-684. Retrieved from http://science.sciencemag.org/content/359/6376/679.abstract
Chung, K., & Deisseroth, K. (2013). CLARITY for mapping the nervous system. Nature Methods, 10, 508. Retrieved from http://dx.doi.org/10.1038/nmeth.2481
Constine, J. (2017). Facebook is building brain-computer interfaces for typing and skin-hearing. TechCrunch. Retrieved from https://techcrunch.com/2017/04/19/facebook-brain-interface/
Cook, S. J., Jarrell, T. A., Brittin, C. A., Wang, Y., Bloniarz, A. E., Yakovlev, M. A., … Emmons, S. W. (2019). Whole-animal connectomes of both Caenorhabditis elegans sexes. Nature, 571(7763), 63–71. https://doi.org/10.1038/s41586-019-1352-7
Cyranoski, D. (2017). China launches brain-imaging factory. Nature, 548(7667), 268–269. https://doi.org/10.1038/548268a
Deadwyler, S., Hampson, R., Sweat, A., Song, D., Chan, R., Opris, I., … Berger, T. (2013). Donor/recipient enhancement of memory in rat hippocampus. Frontiers in Systems Neuroscience. Retrieved from https://www.frontiersin.org/article/10.3389/fnsys.2013.00120
Ding, Y., Vanselow, D. J., Yakovlev, M. A., Katz, S. R., Lin, A. Y., Clark, D. P., … Cheng, K. C. (2019). Computational 3D histological phenotyping of whole zebrafish by X-ray histotomography. ELife, 8, e44898. https://doi.org/10.7554/eLife.44898
Djurfeldt, M., Lundqvist, M., Johansson, C., Rehn, M., Ekeberg, O., & Lansner, A. (2008). Brain-scale simulation of the neocortex on the IBM Blue Gene/L supercomputer. IBM Journal of Research and Development, 52(1.2), 31–41. https://doi.org/10.1147/rd.521.0031
Eberle, A. L., Mikula, S., Schalek, R., Lichtman, J., Tate, M. L. K., & Zeidler, D. (2015). High-resolution, high-throughput imaging with a multibeam scanning electron microscope. Journal of Microscopy, 259(2), 114–120. https://doi.org/10.1111/jmi.12224
Erö, C., Gewaltig, M.-O., Keller, D., & Markram, H. (2018). A Cell Atlas for the Mouse Brain. Frontiers in Neuroinformatics, 12, 84. Retrieved from https://www.frontiersin.org/article/10.3389/fninf.2018.00084
Etherington, D. (2017). Elon Musk’s Neuralink wants to boost the brain to keep up with AI. TechCrunch. Retrieved from techcrunch.com/2017/03/27/elon-musks-neuralink-wants-to-boost-the-brain-to-keep-up-with-ai/
Fact Sheet: BRAIN Initiative. (2013). Retrieved from https://obamawhitehouse.archives.gov/the-press-office/2013/04/02/fact-sheet-brain-initiative
Flesher, S. N., Collinger, J. L., Foldes, S. T., Weiss, J. M., Downey, J. E., Tyler-Kabara, E. C., … Gaunt, R. A. (2016). Intracortical microstimulation of human somatosensory cortex. Science Translational Medicine. Retrieved from http://stm.sciencemag.org/content/early/2016/10/12/scitranslmed.aaf8083.abstract
Gao, R., Asano, S. M., Upadhyayula, S., Pisarev, I., Milkie, D. E., Liu, T.-L., … Betzig, E. (2019). Cortical column and whole-brain imaging with molecular contrast and nanoscale resolution. Science, 363(6424), eaau8302. https://doi.org/10.1126/science.aau8302
Gouwens, N. W., Sorensen, S. A., Berg, J., Lee, C., Jarsky, T., Ting, J., … Koch, C. (2019). Classification of electrophysiological and morphological neuron types in the mouse visual cortex. Nature Neuroscience, 22(7), 1182–1195. https://doi.org/10.1038/s41593-019-0417-0
Graham, B. J., Hildebrand, D. G. C., Kuan, A. T., Maniates-Selvin, J. T., Thomas, L. A., Shanny, B. L., & Lee, W.-C. A. (2019). High-throughput transmission electron microscopy with automated serial sectioning. BioRxiv, 657346. https://doi.org/10.1101/657346
Gunaydin, L. A., Yizhar, O., Berndt, A., Sohal, V. S., Deisseroth, K., & Hegemann, P. (2010). Ultrafast optogenetic control. Nature Neuroscience, 13, 387. Retrieved from http://dx.doi.org/10.1038/nn.2495
Hampson, R. E., Song, D., Robinson, B. S., Fetterhoff, D., Dakos, A. S., Roeder, B. M., … Deadwyler, S. A. (2018). Developing a hippocampal neural prosthetic to facilitate human memory encoding and recall. Journal of Neural Engineering, 15(3), 36014. https://doi.org/10.1088/1741-2552/aaaed7
Han, X., & Boyden, E. S. (2007). Multiple-Color Optical Activation, Silencing, and Desynchronization of Neural Activity, with Single-Spike Temporal Resolution. PLOS ONE, 2(3), e299. Retrieved from https://doi.org/10.1371/journal.pone.0000299
Hatmaker, T. (2017). DARPA awards $65 million to develop the perfect, tiny two-way brain-computer interface. TechCrunch. Retrieved from techcrunch.com/2017/07/10/darpa-nesd-grants-paradromics/
Hayworth, K. J., Peale, D., Januszewski, M., Knott, G. W., Lu, Z., Xu, C. S., & Hess, H. F. (2019). Gas cluster ion beam SEM for imaging of large tissue samples with 10 nm isotropic resolution. Nature Methods. https://doi.org/10.1038/s41592-019-0641-2
Hildebrand, D. G. C., Cicconet, M., Torres, R. M., Choi, W., Quan, T. M., Moon, J., … Engert, F. (2017). Whole-brain serial-section electron microscopy in larval zebrafish. Nature, 545, 345. Retrieved from https://doi.org/10.1038/nature22356
Horton, N. G., Wang, K., Kobat, D., Clark, C. G., Wise, F. W., Schaffer, C. B., & Xu, C. (2013). In vivo three-photon microscopy of subcortical structures within an intact mouse brain. Nature Photonics, 7, 205. Retrieved from https://doi.org/10.1038/nphoton.2012.336
Jiang, L., Stocco, A., Losey, D. M., Abernethy, J. A., Prat, C. S., & Rao, R. P. N. (2018). BrainNet: a multi-person brain-to-brain interface for direct collaboration between brains. ArXiv Preprint ArXiv:1809.08632.
Jordan, J., Ippen, T., Helias, M., Kitayama, I., Sato, M., Igarashi, J., … Kunkel, S. (2018). Extremely Scalable Spiking Neuronal Network Simulation Code: From Laptops to Exascale Computers. Frontiers in Neuroinformatics, 12, 2. Retrieved from https://www.frontiersin.org/article/10.3389/fninf.2018.00002
Jun, J. J., Steinmetz, N. A., Siegle, J. H., Denman, D. J., Bauza, M., Barbarits, B., … Harris, T. D. (2017). Fully integrated silicon probes for high-density recording of neural activity. Nature, 551, 232. Retrieved from https://doi.org/10.1038/nature24636
Kanari, L., Ramaswamy, S., Shi, Y., Morand, S., Meystre, J., Perin, R., … Markram, H. (2019). Objective Morphological Classification of Neocortical Pyramidal Cells. Cerebral Cortex, 29(4), 1719–1735. https://doi.org/10.1093/cercor/bhy339
Kasthuri, N., Hayworth, K. J., Berger, D. R., Schalek, R. L., Conchello, J. A., Knowles-Barley, S., … Lichtman, J. W. (2015). Saturated Reconstruction of a Volume of Neocortex. Cell, 162(3), 648–661. https://doi.org/10.1016/j.cell.2015.06.054
Kato, S., Kaplan, H. S., Schrödel, T., Skora, S., Lindsay, T. H., Yemini, E., … Zimmer, M. (2015). Global Brain Dynamics Embed the Motor Command Sequence of Caenorhabditis elegans. Cell, 163(3), 656–669. https://doi.org/10.1016/j.cell.2015.09.034
Kemen, T., Malloy, M., Thiel, B., Mikula, S., Denk, W., Dellemann, G., & Zeidler, D. (2015). Further advancing the throughput of a multi-beam SEM. In Proc.SPIE (Vol. 9424). Retrieved from https://doi.org/10.1117/12.2188560
Lee, J. H., Daugharthy, E. R., Scheiman, J., Kalhor, R., Yang, J. L., Ferrante, T. C., … Church, G. M. (2014). Highly Multiplexed Subcellular RNA Sequencing in Situ. Science, 343(6177), 1360 LP-1363. https://doi.org/10.1126/science.1250212
Li, P. H., Lindsey, L. F., Januszewski, M., Zheng, Z., Bates, A. S., Taisz, I., … Jain, V. (2019). Automated Reconstruction of a Serial-Section EM Drosophila Brain with Flood-Filling Networks and Local Realignment. BioRxiv, 605634. https://doi.org/10.1101/605634
Lin, M. K., Takahashi, Y. S., Huo, B.-X., Hanada, M., Nagashima, J., Hata, J., … Mitra, P. (2019). A high-throughput neurohistological pipeline for brain-wide mesoscale connectivity mapping of the common marmoset. ELife, 8, e40042. https://doi.org/10.7554/eLife.40042
Liu, J., Fu, T.-M., Cheng, Z., Hong, G., Zhou, T., Jin, L., … Lieber, C. M. (2015). Syringe-injectable electronics. Nature Nanotechnology, 10, 629. Retrieved from http://dx.doi.org/10.1038/nnano.2015.115
Livet, J., Weissman, T. A., Kang, H., Draft, R. W., Lu, J., Bennis, R. A., … Lichtman, J. W. (2007). Transgenic strategies for combinatorial expression of fluorescent proteins in the nervous system. Nature, 450, 56. Retrieved from http://dx.doi.org/10.1038/nature06293
Lomas, N. (2017). Superintelligent AI explains Softbank’s push to raise a $100BN Vision Fund. TechCrunch. Retrieved from https://techcrunch.com/2017/02/27/superintelligent-ai-explains-softbanks-push-to-raise-a-100bn-vision-fund/
Ma, Y., Bao, J., Zhang, Y., Li, Z., Zhou, X., Wan, C., … Xue, T. (2019). Mammalian Near-Infrared Image Vision through Injectable and Self-Powered Retinal Nanoantennae. Cell, 177(2), 243–255.e15. https://doi.org/https://doi.org/10.1016/j.cell.2019.01.038
Mardinly, A. R., Oldenburg, I. A., Pégard, N. C., Sridharan, S., Lyall, E. H., Chesnov, K., … Adesnik, H. (2018). Precise multimodal optical control of neural ensemble activity. Nature Neuroscience, 21(6), 881–893. https://doi.org/10.1038/s41593-018-0139-8
Markram, H. (2006). The Blue Brain Project. Nature Reviews Neuroscience, 7, 153. Retrieved from http://dx.doi.org/10.1038/nrn1848
Markram, H., Muller, E., Ramaswamy, S., Reimann, M. W., Abdellah, M., Sanchez, C. A., … Schürmann, F. (2015). Reconstruction and Simulation of Neocortical Microcircuitry. Cell, 163(2), 456–492. https://doi.org/10.1016/j.cell.2015.09.029
Marx, V. (2013). Neuroscience waves to the crowd. Nature Methods, 10, 1069. Retrieved from http://dx.doi.org/10.1038/nmeth.2695
Milano, G., Luebben, M., Ma, Z., Dunin-Borkowski, R., Boarino, L., Pirri, C. F., … Valov, I. (2018). Self-limited single nanowire systems combining all-in-one memristive and neuromorphic functionalities. Nature Communications, 9(1), 5151. https://doi.org/10.1038/s41467-018-07330-7
Mizutani, R., Saiga, R., Takeuchi, A., Uesugi, K., & Suzuki, Y. (2013). Three-dimensional network of Drosophila brain hemisphere. Journal of Structural Biology, 184(2), 271–279. https://doi.org/https://doi.org/10.1016/j.jsb.2013.08.012
Motta, A., Berning, M., Boergens, K. M., Staffler, B., Beining, M., Loomba, S., … Helmstaedter, M. (2019). Dense connectomic reconstruction in layer 4 of the somatosensory cortex. Science, 366(6469), eaay3134. https://doi.org/10.1126/science.aay3134
Murakami, T. C., Mano, T., Saikawa, S., Horiguchi, S. A., Shigeta, D., Baba, K., … Ueda, H. R. (2018). A three-dimensional single-cell-resolution whole-brain atlas using CUBIC-X expansion microscopy and tissue clearing. Nature Neuroscience, 21(4), 625–637. https://doi.org/10.1038/s41593-018-0109-1
Musk, E. (2019). An integrated brain-machine interface platform with thousands of channels. BioRxiv, 703801. https://doi.org/10.1101/703801
Nemrodov, D., Niemeier, M., Patel, A., & Nestor, A. (2018). The Neural Dynamics of Facial Identity Processing: Insights from EEG-Based Pattern Analysis and Image Reconstruction. Eneuro, 5(1), ENEURO.0358-17.2018. https://doi.org/10.1523/ENEURO.0358-17.2018
Nishiyama, J., Mikuni, T., & Yasuda, R. (2017). Virus-Mediated Genome Editing via Homology-Directed Repair in Mitotic and Postmitotic Cells in Mammalian Brain. Neuron, 96(4), 755–768.e5. https://doi.org/10.1016/j.neuron.2017.10.004
Oizumi, M., Albantakis, L., & Tononi, G. (2014). From the Phenomenology to the Mechanisms of Consciousness: Integrated Information Theory 3.0. PLOS Computational Biology, 10(5), e1003588. Retrieved from https://doi.org/10.1371/journal.pcbi.1003588
Okano, H., Miyawaki, A., & Kasai, K. (2015). Brain/MINDS: brain-mapping project in Japan. Philosophical Transactions of the Royal Society of London. Series B, Biological Sciences, 370(1668). https://doi.org/10.1098/rstb.2014.0310
Pacureanu, A., Maniates-Selvin, J., Kuan, A. T., Thomas, L. A., Chen, C.-L., Cloetens, P., & Lee, W.-C. A. (2019). Dense neuronal reconstruction through X-ray holographic nano-tomography. BioRxiv, 653188. https://doi.org/10.1101/653188
Peng, H., Hawrylycz, M., Roskams, J., Hill, S., Spruston, N., Meijering, E., & Ascoli, G. A. (2015). BigNeuron: Large-Scale 3D Neuron Reconstruction from Optical Microscopy Images. Neuron, 87(2), 252–256. https://doi.org/10.1016/j.neuron.2015.06.036
Peng, H., Zhou, Z., Meijering, E., Zhao, T., Ascoli, G. A., & Hawrylycz, M. (2017). Automatic tracing of ultra-volumes of neuronal images. Nature Methods, 14, 332. Retrieved from https://doi.org/10.1038/nmeth.4233
Poo, M., Du, J., Ip, N. Y., Xiong, Z.-Q., Xu, B., & Tan, T. (2016). China Brain Project: Basic Neuroscience, Brain Diseases, and Brain-Inspired Computing. Neuron, 92(3), 591–596. https://doi.org/10.1016/j.neuron.2016.10.050
Regalado, A. (2017). The Entrepreneur with the $100 Million Plan to Link Brains to Computers. MIT Technology Review. Retrieved from https://www.technologyreview.com/s/603771/the-entrepreneur-with-the-100-million-plan-to-link-brains-to-computers/
Reimann, M. W., Gevaert, M., Shi, Y., Lu, H., Markram, H., & Muller, E. (2019). A null model of the mouse whole-neocortex micro-connectome. Nature Communications, 10(1), 3903. https://doi.org/10.1038/s41467-019-11630-x
Reimann, M. W., Nolte, M., Scolamiero, M., Turner, K., Perin, R., Chindemi, G., … Markram, H. (2017). Cliques of Neurons Bound into Cavities Provide a Missing Link between Structure and Function. Frontiers in Computational Neuroscience. Retrieved from https://www.frontiersin.org/article/10.3389/fncom.2017.00048
Schalek, R., Kasthuri, N., Hayworth, K., Berger, D., Tapia, J., Morgan, J., … Lichtman, J. (2011). Development of High-Throughput, High-Resolution 3D Reconstruction of Large-Volume Biological Tissue Using Automated Tape Collection Ultramicrotomy and Scanning Electron Microscopy. Microscopy and Microanalysis, 17(S2), 966–967. https://doi.org/DOI: 10.1017/S1431927611005708
Schemmel, J., Briiderle, D., Griibl, A., Hock, M., Meier, K., & Millner, S. (2010). A wafer-scale neuromorphic hardware system for large-scale neural modeling. In Proceedings of 2010 IEEE International Symposium on Circuits and Systems (pp. 1947–1950). https://doi.org/10.1109/ISCAS.2010.5536970
Schemmel, J., Kriener, L., Müller, P., & Meier, K. (2017). An accelerated analog neuromorphic hardware system emulating NMDA- and calcium-based non-linear dendrites. In 2017 International Joint Conference on Neural Networks (IJCNN) (pp. 2217–2226). https://doi.org/10.1109/IJCNN.2017.7966124
Seo, D., Neely, R. M., Shen, K., Singhal, U., Alon, E., Rabaey, J. M., … Maharbiz, M. M. (2016). Wireless Recording in the Peripheral Nervous System with Ultrasonic Neural Dust. Neuron, 91(3), 529–539. https://doi.org/10.1016/j.neuron.2016.06.034
Song, D., She, X., Hampson, R. E., Deadwyler, S. A., & Berger, T. W. (2017). Multi-resolution multi-trial sparse classification model for decoding visual memories from hippocampal spikes in human. In 2017 39th Annual International Conference of the IEEE Engineering in Medicine and Biology Society (EMBC) (pp. 1046–1049). https://doi.org/10.1109/EMBC.2017.8037006
Stocco, A., Prat, C. S., Losey, D. M., Cronin, J. A., Wu, J., Abernethy, J. A., & Rao, R. P. N. (2015). Playing 20 Questions with the Mind: Collaborative Problem Solving by Humans Using a Brain-to-Brain Interface. PLOS ONE, 10(9), e0137303. Retrieved from https://doi.org/10.1371/journal.pone.0137303
Suk, H.-J., van Welie, I., Kodandaramaiah, S. B., Allen, B., Forest, C. R., & Boyden, E. S. (2017). Closed-Loop Real-Time Imaging Enables Fully Automated Cell-Targeted Patch-Clamp Neural Recording In Vivo. Neuron, 95(5), 1037–1047.e11. https://doi.org/https://doi.org/10.1016/j.neuron.2017.08.011
Szigeti, B., Gleeson, P., Vella, M., Khayrulin, S., Palyanov, A., Hokanson, J., … Larson, S. (2014). OpenWorm: an open-science approach to modeling Caenorhabditis elegans. Frontiers in Computational Neuroscience. Retrieved from https://www.frontiersin.org/article/10.3389/fncom.2014.00137
Tasic, B., Yao, Z., Graybuck, L. T., Smith, K. A., Nguyen, T. N., Bertagnolli, D., … Zeng, H. (2018). Shared and distinct transcriptomic cell types across neocortical areas. Nature, 563(7729), 72–78. https://doi.org/10.1038/s41586-018-0654-5
Truckenbrodt, S., Maidorn, M., Crzan, D., Wildhagen, H., Kabatas, S., & Rizzoli, S. O. (2018). X10 expansion microscopy enables 25-nm resolution on conventional microscopes. EMBO Reports, 19(9), e45836. https://doi.org/10.15252/embr.201845836
Ukani, N. H., Yeh, C.-H., Tomkins, A., Zhou, Y., Florescu, D., Ortiz, C. L., … Lazar, A. A. (2019). The Fruit Fly Brain Observatory: From Structure to Function. BioRxiv, 580290. https://doi.org/10.1101/580290
Wang, X., Allen, W. E., Wright, M. A., Sylwestrak, E. L., Samusik, N., Vesuna, S., … Deisseroth, K. (2018). Three-dimensional intact-tissue sequencing of single-cell transcriptional states. Science, 361(6400), eaat5691. https://doi.org/10.1126/science.aat5691
Winnubst, J., Bas, E., Ferreira, T. A., Wu, Z., Economo, M. N., Edson, P., … Chandrashekar, J. (2019). Reconstruction of 1,000 Projection Neurons Reveals New Cell Types and Organization of Long-Range Connectivity in the Mouse Brain. Cell, 179(1), 268–281.e13. https://doi.org/10.1016/j.cell.2019.07.042
Wyss Institute launches ReadCoor to commercialize 3D in situ gene sequencing technology. (2016). Wyss Institute. Retrieved from https://wyss.harvard.edu/news/wyss-institute-launches-readcoor-to-commercialize-3d-in-situ-gene-sequencing-technology/
Yang, X., Zhou, T., Zwang, T. J., Hong, G., Zhao, Y., Viveros, R. D., … Lieber, C. M. (2019). Bioinspired neuron-like electronics. Nature Materials, 18(5), 510–517. https://doi.org/10.1038/s41563-019-0292-9
Yin, W., Brittain, D., Borseth, J., Scott, M. E., Williams, D., Perkins, J., … da Costa, N. M. (2019). A Petascale Automated Imaging Pipeline for Mapping Neuronal Circuits with High-throughput Transmission Electron Microscopy. BioRxiv, 791889. https://doi.org/10.1101/791889
Zhang, H.-T., Zuo, F., Li, F., Chan, H., Wu, Q., Zhang, Z., … Ramanathan, S. (2019). Perovskite nickelates as bio-electronic interfaces. Nature Communications, 10(1), 1651. https://doi.org/10.1038/s41467-019-09660-6
Zhao, W., Garcia-Oscos, F., Dinh, D., & Roberts, T. F. (2019). Inception of memories that guide vocal learning in the songbird. Science, 366(6461), 83 LP-89. https://doi.org/10.1126/science.aaw4226
Zheng, Z., Lauritzen, J. S., Perlman, E., Robinson, C. G., Nichols, M., Milkie, D., … Bock, D. D. (2018). A Complete Electron Microscopy Volume of the Brain of Adult Drosophila melanogaster. Cell, 174(3), 730–743.e22. https://doi.org/10.1016/j.cell.2018.06.019
One Comment
Comments are closed.