Overview
Let V be a set of graph vertices embedded in some manifold M and E be a set of graph edges embedded in ℝ. Since manifolds are locally homeomorphic to Euclidean space, every open set a ∈ M can be mapped to an open set b ∈ ℝ by a homeomorphism f. Likewise, for every f there exists an inverse f -1 which maps from ℝ to M.
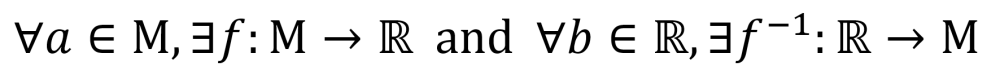
The distance L between vertices u and v will be defined by the Euclidean metric in ℝ and the vertices will be defined in terms of Euclidean coordinates.
Example: a Graph on a 2-Manifold
Consider a 2-manifold M embedded in ℝ3 and defined by a multivariable function h(x,y) with open boundaries (-5,5), (-5,5), and (-1,1) for x, y, and z respectively.

Let V be a set of graph vertices be embedded in M. Every vertex v ∈ V is defined as a point v ∈ M and every edge vivj ∈ E is defined as a parameterized curve s ⊆ M for which the endpoints of s are the vertices vi ∈ V and vj ∈ V.

Figure 1 (A) Perspective view of the 2-manifold M and graph G. (B) Since M is embedded in ℝ3 and the vertices v ⊆ M, the Euclidean coordinates for each vertex are listed as ordered triples. The distance L between any two points or graph vertices u and v is given by the shortest curve s which can be parameterized on M from u to v. The distance L must satisfy the axioms of a metric on M. (C) Adjacency matrix for G. (D) M and G as a projection on ℝ2 (a “top view”). (E) Perspective view of the graph G embedded in ℝ3 without M.
In this example, five curves corresponding with the five edges of G will be parameterized. The following steps were used to generate these parameterizations in MATLAB.
- Considering only ℝ2 (a “top view”), the equations of line segments between vi and vj were found by determining the slopes and y-intercepts from the x and y coordinates of vi and vj. This equation took the form y = mx + b.
- The commands x=[-5:0.1:5] and y=[-5:0.1:5] were inputted to generate vectors of data for x and y.
- The equation of the surface, h(x + y) = sin(x + y) was created.
- MATLAB’s “fit” tool was used to fit parameterized curves to the surface. The command took the form fit(mx + b, h(x, mx + b), ‘sin1’).
- The symbolic vector equations were generated by setting each x equal to m-1t, each y equal to t plus the y-intercept, and each z equal to sin(k1t + k2) where the constants were provided by the “fit” tool from the previous step.
- The resulting functions were visualized along with the surface h. Since they were still phase shifted relative to the surface, these shifts were manually corrected. It should be noted that this step would not be effective with manifolds that cannot be easily visualized, so for more abstract applications, improvements to this procedure will be necessary.
- The domain of t was computed. The final vector equations are recorded below.

From these parametric equations, arc lengths can be computed by using the arc length formula in ℝ3. Here, the parameterization and arc length formula are utilized for illustrative purposes, but it should be noted that other methods for determining arc length might be superior in many cases.

For instance, rather than fitting parametric curves onto a 2-manifold, the graph edges might be described in terms of numerous short tangent line segments. Together, these segments would approximate the appropriate parametric curves. By summing the lengths of the small segments, arc lengths for graph edges could be numerically approximated. Furthermore, this type of numerical technique could be extended to manifolds embedded in non-Euclidean spaces (i.e. complex manifolds, manifolds embedded in elliptic spaces, etc.) by defining an appropriate metric for the given space.

Table 1 Arc lengths for the edges of G embedded in M. These values were computed from the parametric curves describing the edges.
Now that G has been characterized in M, the network can be locally mapped into ℝ2. There exist many possible sets of homeomorphisms which could map open sets of points in M (which together contain G) to ℝ2. In the broader context, depending on the modeling application, different sets of maps might be chosen. Here, the mappings will project to ℝ2 along the z-axis. The open sets p1, p2, p3, and p4 centered on each vertex v ∈ V will have arbitrarily defined radii r1,2,3,4 = 5 on the manifold M and project to open sets q1, q2, q3, and q4 in ℝ2. In this case, the radii of these open sets do not affect the mappings so long as the open sets cover the graph G and the following conditions hold.

As such, the Euclidean distance metric on ℝ2 (above) can be applied to globally change the edge lengths from the lengths given in table 1 and the middle column of table 2 to those given in the rightmost column of table 2.

Table 2 Lengths for the edges of G projected into ℝ2 compared with the values from table 1. The lengths in ℝ2 were computed using the Euclidean distance metric.
Applications
While the example provided in this text represents a somewhat arbitrary situation, it illustrates a general process which might be extended for many applications. Many phenomena in nature reflect particular geometric properties which could be amenable to modeling with manifolds. When the given geometries influence the properties of real-world networks, my technique may provide a useful model.
For instance, consider the topographic maps found in parts of sensory (i.e. retinotopic, somatotopic, tonotopic, etc.) and motor cortices. Neurons in these regions are spatially arrayed in a manner which corresponds to the physical characteristics of sensory data. In the case of retinotopy, the spatial coordinates of an observed image have a direct correspondence to the spatial coordinates on the retina. The coordinates on the retina in turn have a direct correspondence to coordinates in the primary visual cortex (V1). Higher cortical areas have more complicated retinotopic maps in which adjacent subsets of the visual field are not always adjacent to each other in the given cortical region. The curved surface of the retina could be described as a 2-manifold which takes in deformed versions of the final images that the brain reconstructs. In addition, the cortical folds in V1 could be described by another 2-manifold. Since the final percept (what a person consciously “sees”) is reconstructed into an image that can be considered as roughly Euclidean, the local Euclidean properties of manifolds can be applied to this situation.
By fitting a parameterized surface to imaging data which show the folded geometry in V1 (or other topographic sensory or motor areas) and applying a graphical model with mesoscale (or higher) resolution of connectivity among regions within this cortex, global morphological changes which influence the final percept could be modeled. This may be useful for gleaning insights about congenital brain abnormalities and traumatic brain injuries. In addition, this method could be valuable for developmental neurobiology. Surfaces could be fitted to cortical geometries at multiple time-points in development and relationships between network structure, manifold geometry, and fetal brain activity in higher cortical regions (or their developing equivalents) could be analyzed. (Thomason et al. recently demonstrated a technique for fMRI in utero). Of course, it would be difficult to determine the types of percepts that a fetus might generate, so a different mathematical space may need to be constructed based upon functional activity in the developing downstream visual processing regions.
Here, only 2-manifolds are discussed. However, many other classes of manifolds could be amenable to modeling in this way. The range of applications could be greatly expanded by changing the requirement for a manifold to be locally Euclidean to a more general condition. Such manifolds would need to be locally homeomorphic to some chosen coordinate space, the particular space depending on the application (i.e. the complex plane, elliptic spaces, polynomial spaces, etc.) In this scenario, 2-manifolds and 3-manifolds which are locally homeomorphic to coordinate systems defined by other deformed surfaces and volumes would probably be particularly applicable. By constructing manifolds which are locally homeomorphic to other manifolds and computing how embedded networks are transformed across these manifolds, a myriad of potential phenomena could be analyzed. This method may open new possibilities for modeling of biological, technological, social, physical, and other real-world networks which are influenced by geometric factors.
This text represents one of my first forays into developing new mathematical tools. Given that I come from a biological background, it may appear quite rough to more mathematically-experienced individuals. However, I feel that this technique has potential to develop further and be applied to problems in science and engineering. I would welcome any constructive critiques so long as they are presented in a respectful manner.