PDF version: A Guide to CRISPR-Cas Nucleases by Logan Thrasher Collins
Many different types of CRISPR-Cas nucleases possess biotechnological relevance. For a newcomer, the menagerie of Cas proteins may seem overwhelming. It can be challenging to decide which type of CRISPR system to employ in one’s research. To help address this issue, I compiled these notes. While my guide is certainly not comprehensive, it still covers a wide swath of important Cas proteins and may prove valuable as a starting point for those interested in getting a sense of the field. One should be aware that the field of CRISPR technology is moving rapidly, so some of the nucleases described here might eventually be superseded by newly discovered and/or newly engineered Cas proteins. I would also like to mention that since these notes are specifically focused on types of Cas proteins, I have omitted direct explanations of some important CRISPR technologies such as base editors, prime editors, and dead Cas systems. I also have not directly explained important CRISPR-related concepts such as non-homologous end joining (NHEJ), homology-directed repair (HDR), and adeno-associated virus (AAV) vectors. I encourage the reader to look elsewhere to learn about these subjects since they are vital for having a strong understanding of CRISPR biotechnology. I hope that you enjoy reading my notes and find them useful for your own scientific endeavors!
SpCas9

SpCas9 represents one of the first discovered and most commonly used CRISPR-Cas proteins.1 It comes from Streptococcus pyogenes, a gram-positive bacterial pathogen. SpCas9 employs two nuclease domains to make blunt double-stranded cuts in DNA: the HNH domain for cutting the strand which pairs with the gRNA and the RuvC domain for cutting the other strand. The protospacer adjacent motif (PAM) of SpCas9 has the sequence 5’-NGG-3’, which limits the target sites that the nuclease can find. Though wild-type (WT) SpCas9 possesses a problematic level of off-target activity, several mutant variants of the enzyme have been engineered which give it much more precision.2,3 As some examples, a few of these (but not all of them) include eSpCas9-HF, eSpCas9(1.1), and HypaCas9. The eSpCas9-HF and eSpCas9(1.1) enzymes maintain robust on-target cleavage while reducing off-target effects.3 The HypaCas9 enzyme has similar properties, but with even less off-target effects.2
SaCas9

At 1053 amino acids in length, SaCas9 is significantly smaller than SpCas9 (which is 1368 amino acids long).4 SaCas9 can be used in mammalian cells, employs NNGRRT PAM sites (R is A or G), and uses RuvC and HNH domains for cutting. But without further engineering, SaCas9 has lower target specificity even than SpCas9. Fortunately, mutant versions of SaCas9 which exhibit improved targeting accuracy have been developed. Tan et al. engineered SaCas9-HF, a version of the protein which has much less off-target activity relative to the WT SaCas9 and retains its on-target activity.4 With such improvements, SaCas9-HF can serve as a useful alternative to SpCas9.
LbCas12a
The LbCas12a enzyme makes staggered cuts using a single RuvC domain (and no HNH domain), uses T-rich PAM sites, and catalyzes its own crRNA maturation.5 LbCas12a comes from Lachnospiraceae bacterium ND2006. LbCas12a has another remarkable property: the binding and cleavage of target dsDNA activates a separate part of the protein which nonspecifically cleaves any ssDNA in its vicinity. This nonspecific trans-cleavage activity is thought to occur as a result of a conformational change in the LbCas12a protein which exposes its RuvC domain for broader ssDNA attack after binding to target dsDNA.6 It should be noted that other type-V Cas proteins including AsCas12a (see corresponding section), FnCas12a (from the bacterium Francisella novicida), and AaCas12b (from the bacterium Alicyclobacillus acidoterrestris) have been shown to exhibit the same capabilities.5 There furthermore exist many RNA-guided RNA-targeting Cas proteins which possess the same types of abilities.7 There are likely many other type-V Cas proteins with these capabilities as well. The activation of type-V Cas proteins to perform indiscriminate ssDNA cleavage after exposure to target dsDNA has been exploited as a target-induced signal amplification method to develop novel molecular diagnostics.6
AsCas12a

The AsCas12a protein (also called Cpf1) is derived from Acidaminococcus sp.,8 which are a group of anaerobic gram-negative bacteria. The protein exhibits several distinctive features compared to Cas9. AsCas12a utilizes a T-rich PAM site, unlike Cas9’s G-rich PAM. This is useful since it expands the possible targets for CRISPR. In particular, the T-rich PAM of AsCas12a can be useful when dealing with organisms that have AT-rich genomes such as Plasmodium falciparum. The naturally occurring form of AsCas12a does not require a tracrRNA, instead its CRISPR arrays are processed into just crRNAs, which serve to complete the functional AsCas12a-crRNA complex. Rather than creating blunt ends, AsCas12a makes staggered cuts with 4-5 nucleotide 5’ overhangs. This is useful since it increases the precision of non-homologous end joining (NHEJ) repair and allows insertion of DNA sequences at a chosen cut site with a desired orientation as specified by the base pairing of the insert with the overhang sequences. In addition, the AsCas12a protein employs a single RuvC domain to make its staggered cuts and does not have an HNH domain. AsCas12a has a lower tolerance for gRNA-target mismatches9 compared to SpCas9 and therefore demonstrates greater targeting specificity. As a result, AsCas12a shows fewer off-target effects overall. But it also has a lower editing efficiency compared to Cas9 proteins, which means that less cells receive any edits upon introduction of the AsCas12a. As described with LbCas12a, the AsCas12a protein also can carry out nonspecific ssDNA cleavage after it cuts to its target dsDNA.
AsCas12a ultra
WT AsCas12a possesses high targeting specificity, low off-target effects, and makes 5’ overhangs which facilitate correct insert orientation (see the section on AsCas12a). These properties represent desirable qualities for therapeutic gene editing, but the low editing efficiency of AsCas12a limits its therapeutic potential. Because of this, Zhang et al. (in a collaboration between Editas and Integrated DNA Technologies) developed an engineered version of the protein which was dubbed AsCas12a ultra.9 This AsCas12a ultra protein was created using directed evolution in bacteria. It has two point mutations relative to WT AsCas12a, M537R and F870L. These mutations grant the AsCas12a ultra extremely high editing efficiency while maintaining the protein’s low level of off-target effects. For a variety of target sites, Zhang et al. demonstrated nearly 100% editing efficiency in HSPCs, iPSCs, T cells, and NK cells using AsCas12a ultra. They also showed 93% efficiency for simultaneous disruption of three genes in T cells. When performing knock-in edits, Zhang et al. achieved efficiencies of 60% in T cells, 50% in NK cells, and 30% in HSPCs. These impressive numbers illustrate the utility of AsCas12a ultra as a broadly applicable tool for therapeutic gene editing.
AsCas12f1
The AsCas12f1 protein consists of only 422 amino acids, making it one of the smallest Cas proteins known.10 It comes from a type of gram-positive iron-oxidizing bacteria called Acidibacillus sulfuroxidans. AsCas12f1 makes staggered double-stranded breaks in target DNA and recognizes 5’ T-rich PAMs. Even with minimal engineering (just the construction of gRNA from combining its tracrRNA and mature crRNA), Wu et al. showed that AsCas12f1 exhibits usable levels of activity in mammalian cells.10 When expressed directly in mammalian cells via a plasmid, the protein achieved a maximum indel efficiency of 32.8%. When delivered to mammalian cells by AAV-DJ, the maximum indel efficiency was 11.5%. The AsCas12f1 protein possesses considerable promise as a compact therapeutic gene editing tool.
Kim et al.’s engineered Un1Cas12f
At 529 amino acids in length, the Un1Cas12f nuclease represents one of the smallest Cas proteins yet discovered.11 This is useful since the small size of Un1Cas12f’s gene allows it to easily fit within AAV vectors. It comes from an uncultured archaeon and is classified as a type-V CRISPR nuclease, which utilize a C-terminal RuvC domain and do not possess an HNH domain. Though the original Un1Cas12f-gRNA complex has very low editing efficiency in eukaryotic cells, Kim et al. were able to intensively engineer the gRNA using a rational design strategy and achieve an 867-fold improvement of indel frequency in mammalian cells.12 They also showed that the Un1Cas12f gene and gRNA gene could be delivered to the cells using AAVs. Because of its small size, Un1Cas12f may serve as an excellent scaffold for creating base editors and prime editors which fit inside of AAVs.
CasMINI
The CasMINI protein is another engineered CRISPR nuclease derived from Cas12f,13 which comes from an uncultured archaeon. This Cas12f is the same as the Un1Cas12f used by Kim et al.12 Since Cas12f has little to no editing activity in mammalian cells, Xu et al. used rational design to optimize the associated gRNA and employed directed evolution to optimize the protein itself.13 CasMINI, a 529 amino acid protein, was the end result of these approaches. When CasMINI was modified to make dCasMINI-VPR (the VPR is a protein fusion which activates certain genes), it performed with comparable efficiency relative to the commonly used dLbCas12a-VPR. In some cases, dCasMINI-VPR actually outperformed dLbCas12a-VPR. When dCasMINI was modified by fusing base editor (ABE) domains at its N-terminus, the dCasMINI-ABE constructs performed base editing at comparable efficiency relative to dLbCas12a-ABE proteins. Because of their small sizes, the genes encoding the dCasMINI-ABE designs could easily fit into AAV vectors, though Xu et al. did not test this in their paper. Furthermore, even the genes encoding CasMINI prime editors should fit into AAV vectors. It should be noted that the most efficient dCasMINI-ABE base editing occurred in a narrow window precisely 3-4 bp downstream of the PAM site. When CasMINI was tested for its ability to perform gene editing by making indels, it showed significantly improved activity over Cas12f, though the editing efficiencies were still fairly low at around 5-10%.
Cas12j

The Cas12j enzyme, also known as CasΦ, comes from the genomes of huge bacteriophages of the Biggiephage clade.14 This is remarkable since CRISPR systems have usually been found in bacteria and archaea rather than viruses (though the prevalence of such machinery in viruses is perhaps underestimated). It has been hypothesized that Biggiephages use Cas12j to cut the DNA of other competing bacteriophages. There exist subtypes of Cas12j such as Cas12j-1, Cas12j-2, and Cas12j-3. All of the Cas12j nucleases are small at between 700 and 800 amino acids in length. The Cas12j nuclease cuts target dsDNA using a single C-terminal RuvC domain. Cas12j’s RuvC domain has a small amount of homology to the TnpB protein superfamily from which type-V Cas proteins evolved, yet it still shares <7% amino acid identity overall with type-V Cas proteins. Cas12j is most closely related to a type of TnpB group which is distinct from the type-V enzymes. The Cas12j nuclease catalyzes its own crRNA maturation using its RuvC domain (similar to the type-V nucleases). Unlike the type-V Cas proteins, Cas12j uses the same active site for both its RuvC cleavage of target DNA and its RuvC processing of the crRNA. It employs T-rich PAM sites which have fairly minimal target requirements. For example, the PAM of the Cas12j-2 subtype is 5’-TBN-3’ (B = G, T, or C). These minimal requirements give Cas12j expanded target recognition capabilities compared to other Cas proteins. Cas12j is active in vitro as well as within bacterial, human, and plant cells. Cas12j-2 (with a gRNA) has been observed to edit up to 33% of HEK293 cells. Though this may sound somewhat low, it represents an editing efficiency comparable to that initially reported for Cas9.
LwaCas13a
The LwaCas13a protein is a type-VI CRISPR nuclease and it cleaves RNA rather than DNA.15 It represents one of the most active types of RNA-guided RNA-targeting Cas proteins. LwaCas13a catalyzes the maturation of its own crRNA. The enzyme comes from Leptotrichia wadei, a type of anaerobic gram-negative bacteria found in saliva. LwaCas13a has demonstrated around 50%-80% knockdown of target RNAs in mammalian and plant cells. This is similar to the knockdown efficiencies of shRNAs, but LwaCas13a shows much lower off-target effects. When converted into dLwaCas13a, the protein can act as an RNA imaging tool. It has also been reported to have strong potential for therapeutics as well. One of the most important emerging applications of LwaCas13a (and similar Cas proteins) is that they can be used in diagnostics for infectious diseases.7 To do this, the LwaCas13a gRNA can be designed to target an RNA sequence from a desired pathogen. LwaCas13a can then be mixed with a short reporter RNA oligonucleotide which has a fluorophore at one end and a quencher at the other (the fluorophore is quenched by its close proximity to the quencher). If the target pathogen RNA is introduced, LwaCas13a will cleave said target RNA as well as activate nonspecific trans-cleavage activity (see section on LbCas12a), leading to cleavage of the reporter oligonucleotides. When the reporter oligonucleotides are cleaved, the fluorophore is released from the quencher, resulting in observable fluorescence. It should be noted that many CRISPR-based diagnostics require some form of target nucleic acid amplification step to increase signal prior to the usage of a Cas protein like LwaCas13a, though ways to mitigate this limitation are undergoing rapid development.16
Cas13bt
Kannan et al. identified Cas13bt1 and Cas13bt3 as useful RNA-targeting CRISPR nucleases since Cas13bt has some activity in human cells.17 Cas13bt1 and Cas13bt3 are small at just 804 amino acids and 775 amino acids respectively. It should be noted that Cas13bt also exhibits nonspecific nonspecific trans-cleavage activity (see section on LbCas12a) after cleaving its RNA target, which may allow its usage in diagnostics. Kannan et al. took advantage of the small sizes of Cas13bt1 and Cas13bt3 to develop compact RNA base editors. They fused an ADAR2 hyperactive adenosine deaminase catalytic domain onto dCas13bt1 and dCas13bt3. The resulting constructs were respectively named REPAIR.t1 and REPAIR.t3 and were shown to facilitate adenosine to inosine conversion in target RNAs. They also fused an ADAR2dd cytidine deaminase domain (which was itself created through directed evolution) onto dCas13bt1 and dCas13bt3. The resulting constructs were respectively named RESCUE.t1 and RESCUE.t3 and were shown to facilitate conversion of cytosine to uracil in target RNAs. Due to the small sizes of Cas13bt enzymes, all of these RNA base editors were small enough to fit inside of AAV vectors even alongside gRNA encoding sequences. The authors demonstrated successful AAV-mediated delivery to cells, but the editing efficiencies were low, so further optimization will likely be necessary.
CasX
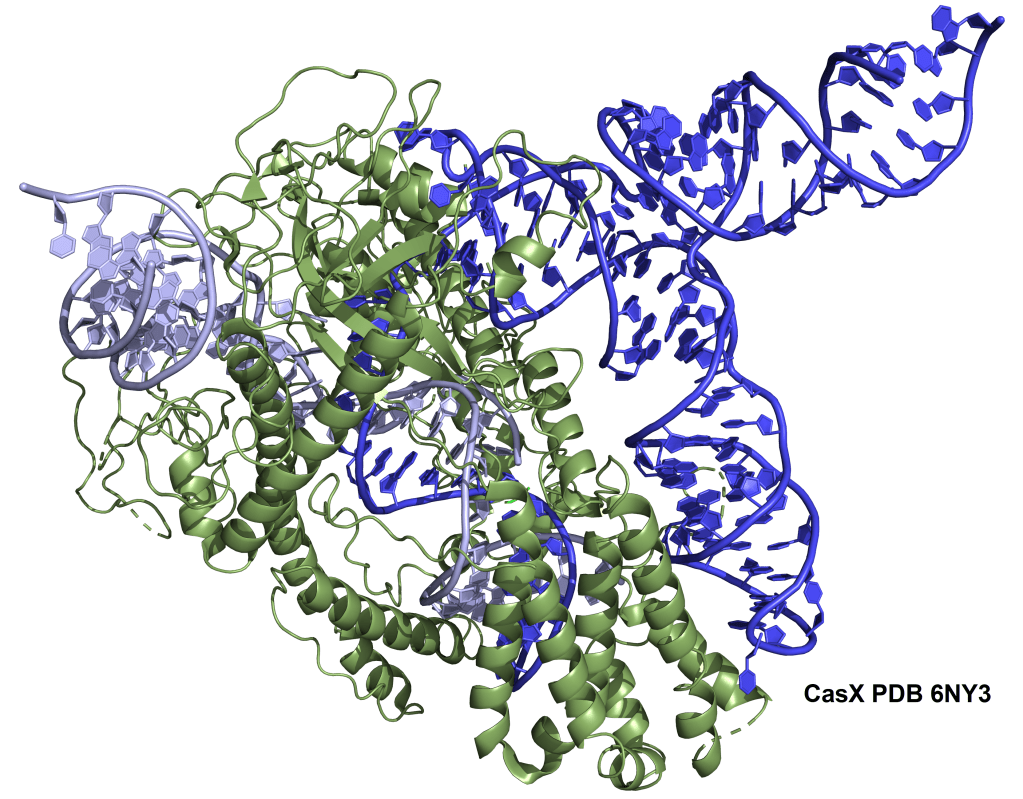
The CasX nuclease represents a distinct type of Cas protein which does not share much sequence similarity with other types of CRISPR enzymes except for a RuvC domain.18 It is an RNA-guided DNA-targeting endonuclease which has minimal nonspecific trans-cleavage activity. Using its single RuvC domain, CasX creates staggered cuts (with about 10 nucleotide overhangs) in dsDNA complementary to its gRNA and adjacent to its TTCN PAM sites. CasX nucleases are <1000 amino acids in length, which is smaller than Cas9 and Cas12a. This could be useful for AAV-mediated delivery of CasX systems. There are different subtypes of CasX which come from different bacteria. Two of the known subtypes are DpbCasX (from Deltaproteobacteria) and PlmCasX (from Planctomycetes). DpbCasx can act in human cells, though it shows limited gene editing efficiency. PlmCasX generally has better efficiency at performing in human cells and can often achieve targeted disruption of genes in around a third of transfected cells. While this level of disruption is still modest, it is similar to the levels originally found with WT Cas9 enzymes before they were optimized for gene editing.
Un1Cas12f (previously known as Cas14a)
The Cas12f proteins represent a class of small CRISPR nucleases (400-700 amino acids in length) that are capable of RNA-guided cleavage of ssDNA or dsDNA depending on whether the gRNA or crRNA includes a PAM. They employ a RuvC domain for cleavage and do not possess an HNH domain. There are various subtypes of Cas12f, but Un1Cas12f (previously Cas14a1) has been studied in the most detail. Un1Cas12f was first reported to selectively cleave ssDNA and not dsDNA.19 It was also initially reported to not require a PAM site for targeting. Without the constraint of needing a PAM site for targeting, Un1Cas12f has broader possibilities for which ssDNA sequences can be targeted. However, later research revealed that Un1Cas12f can cleave dsDNA when a 5’ T-rich PAM sequence is included in the gRNA or crRNA.20 As with many other types of Cas proteins, Un1Cas12f exhibits nonspecific nonspecific trans-cleavage activity of dsDNA (see section on LbCas12a) after cleaving its target DNA, which grants it utility as a component of diagnostics.
DiCas7-11
The Cas7-11 protein is an RNA-guided RNA-targeting CRISPR nuclease.21 It is named Cas7-11 because it arose evolutionarily from a fusion of a protein known as Cas7 with a protein known as Cas11. The DiCas7-11 enzyme comes from the gram-negative sulfate-reducing bacteria Desulfonema ishimotonii (there also exist similar types of Cas7-11 from other species). An important advantage of DiCas7-11 is that it does not have a toxic effect on host cells (bacterial or mammalian). By comparison, RNA knockdown technologies including shRNA, LwaCas13a, PspCas13b, and RfxCas13d typically cause around 30-50% host cell death. DiCas7-11 shows similar knockdown efficiencies compared to these other RNA knockdown technologies while demonstrating no detectable cellular toxicity. Unfortunately, DiCas7-11 is also fairly large at 1602 amino acids, making it difficult to package into AAV vectors. One more application of Cas7-11 is RNA editing. The creation of a dDiCas7-11 fused to a base editor domain has enabled RNA editing in mammalian cells.
References:
3D structure images were created using PyMol.
(1) Anders, C.; Niewoehner, O.; Duerst, A.; Jinek, M. Structural Basis of PAM-Dependent Target DNA Recognition by the Cas9 Endonuclease. Nature 2014, 513 (7519), 569–573. https://doi.org/10.1038/nature13579.
(2) Chen, J. S.; Dagdas, Y. S.; Kleinstiver, B. P.; Welch, M. M.; Sousa, A. A.; Harrington, L. B.; Sternberg, S. H.; Joung, J. K.; Yildiz, A.; Doudna, J. A. Enhanced Proofreading Governs CRISPR–Cas9 Targeting Accuracy. Nature 2017, 550 (7676), 407–410. https://doi.org/10.1038/nature24268.
(3) M., S. I.; Linyi, G.; Bernd, Z.; A., S. D.; X., Y. W.; Feng, Z. Rationally Engineered Cas9 Nucleases with Improved Specificity. Science (80-. ). 2016, 351 (6268), 84–88. https://doi.org/10.1126/science.aad5227.
(4) Tan, Y.; Chu, A. H. Y.; Bao, S.; Hoang, D. A.; Kebede, F. T.; Xiong, W.; Ji, M.; Shi, J.; Zheng, Z. Rationally Engineered Staphylococcus Aureus Cas9 Nucleases with High Genome-Wide Specificity. Proc. Natl. Acad. Sci. 2019, 116 (42), 20969 LP – 20976. https://doi.org/10.1073/pnas.1906843116.
(5) S., C. J.; Enbo, M.; B., H. L.; Maria, D. C.; Xinran, T.; M., P. J.; A., D. J. CRISPR-Cas12a Target Binding Unleashes Indiscriminate Single-Stranded DNase Activity. Science (80-. ). 2018, 360 (6387), 436–439. https://doi.org/10.1126/science.aar6245.
(6) Nalefski, E. A.; Patel, N.; Leung, P. J. Y.; Islam, Z.; Kooistra, R. M.; Parikh, I.; Marion, E.; Knott, G. J.; Doudna, J. A.; Le Ny, A.-L. M.; Madan, D. Kinetic Analysis of Cas12a and Cas13a RNA-Guided Nucleases for Development of Improved CRISPR-Based Diagnostics. iScience 2021, 24 (9), 102996. https://doi.org/https://doi.org/10.1016/j.isci.2021.102996.
(7) Kellner, M. J.; Koob, J. G.; Gootenberg, J. S.; Abudayyeh, O. O.; Zhang, F. SHERLOCK: Nucleic Acid Detection with CRISPR Nucleases. Nat. Protoc. 2019, 14 (10), 2986–3012. https://doi.org/10.1038/s41596-019-0210-2.
(8) Zetsche, B.; Gootenberg, J. S.; Abudayyeh, O. O.; Slaymaker, I. M.; Makarova, K. S.; Essletzbichler, P.; Volz, S. E.; Joung, J.; van der Oost, J.; Regev, A.; Koonin, E. V.; Zhang, F. Cpf1 Is a Single RNA-Guided Endonuclease of a Class 2 CRISPR-Cas System. Cell 2015, 163 (3), 759–771. https://doi.org/https://doi.org/10.1016/j.cell.2015.09.038.
(9) Zhang, L.; Zuris, J. A.; Viswanathan, R.; Edelstein, J. N.; Turk, R.; Thommandru, B.; Rube, H. T.; Glenn, S. E.; Collingwood, M. A.; Bode, N. M.; Beaudoin, S. F.; Lele, S.; Scott, S. N.; Wasko, K. M.; Sexton, S.; Borges, C. M.; Schubert, M. S.; Kurgan, G. L.; et al. AsCas12a Ultra Nuclease Facilitates the Rapid Generation of Therapeutic Cell Medicines. Nat. Commun. 2021, 12 (1), 3908. https://doi.org/10.1038/s41467-021-24017-8.
(10) Wu, Z.; Zhang, Y.; Yu, H.; Pan, D.; Wang, Y.; Wang, Y.; Li, F.; Liu, C.; Nan, H.; Chen, W.; Ji, Q. Programmed Genome Editing by a Miniature CRISPR-Cas12f Nuclease. Nat. Chem. Biol. 2021. https://doi.org/10.1038/s41589-021-00868-6.
(11) Okano, K.; Sato, Y.; Hizume, T.; Honda, K. Genome Editing by Miniature CRISPR/Cas12f1 Enzyme in Escherichia Coli. J. Biosci. Bioeng. 2021, 132 (2), 120–124. https://doi.org/https://doi.org/10.1016/j.jbiosc.2021.04.009.
(12) Kim, D. Y.; Lee, J. M.; Moon, S. Bin; Chin, H. J.; Park, S.; Lim, Y.; Kim, D.; Koo, T.; Ko, J.-H.; Kim, Y.-S. Efficient CRISPR Editing with a Hypercompact Cas12f1 and Engineered Guide RNAs Delivered by Adeno-Associated Virus. Nat. Biotechnol. 2021. https://doi.org/10.1038/s41587-021-01009-z.
(13) Xu, X.; Chemparathy, A.; Zeng, L.; Kempton, H. R.; Shang, S.; Nakamura, M.; Qi, L. S. Engineered Miniature CRISPR-Cas System for Mammalian Genome Regulation and Editing. Mol. Cell 2021. https://doi.org/https://doi.org/10.1016/j.molcel.2021.08.008.
(14) Patrick, P.; Basem, A.-S.; Ezra, B.-R.; A., T. C.; Zheng, L.; F., C. B.; J., K. G.; E., J. S.; F., B. J.; A., D. J. CRISPR-CasΦ from Huge Phages Is a Hypercompact Genome Editor. Science (80-. ). 2020, 369 (6501), 333–337. https://doi.org/10.1126/science.abb1400.
(15) Abudayyeh, O. O.; Gootenberg, J. S.; Essletzbichler, P.; Han, S.; Joung, J.; Belanto, J. J.; Verdine, V.; Cox, D. B. T.; Kellner, M. J.; Regev, A.; Lander, E. S.; Voytas, D. F.; Ting, A. Y.; Zhang, F. RNA Targeting with CRISPR–Cas13. Nature 2017, 550 (7675), 280–284. https://doi.org/10.1038/nature24049.
(16) Kaminski, M. M.; Abudayyeh, O. O.; Gootenberg, J. S.; Zhang, F.; Collins, J. J. CRISPR-Based Diagnostics. Nat. Biomed. Eng. 2021, 5 (7), 643–656. https://doi.org/10.1038/s41551-021-00760-7.
(17) Kannan, S.; Altae-Tran, H.; Jin, X.; Madigan, V. J.; Oshiro, R.; Makarova, K. S.; Koonin, E. V; Zhang, F. Compact RNA Editors with Small Cas13 Proteins. Nat. Biotechnol. 2021. https://doi.org/10.1038/s41587-021-01030-2.
(18) Liu, J.-J.; Orlova, N.; Oakes, B. L.; Ma, E.; Spinner, H. B.; Baney, K. L. M.; Chuck, J.; Tan, D.; Knott, G. J.; Harrington, L. B.; Al-Shayeb, B.; Wagner, A.; Brötzmann, J.; Staahl, B. T.; Taylor, K. L.; Desmarais, J.; Nogales, E.; Doudna, J. A. CasX Enzymes Comprise a Distinct Family of RNA-Guided Genome Editors. Nature 2019, 566 (7743), 218–223. https://doi.org/10.1038/s41586-019-0908-x.
(19) B., H. L.; David, B.; S., C. J.; David, P.-E.; Enbo, M.; P., W. I.; C., C. J.; C., K. N.; F., B. J.; A., D. J. Programmed DNA Destruction by Miniature CRISPR-Cas14 Enzymes. Science (80-. ). 2018, 362 (6416), 839–842. https://doi.org/10.1126/science.aav4294.
(20) Karvelis, T.; Bigelyte, G.; Young, J. K.; Hou, Z.; Zedaveinyte, R.; Budre, K.; Paulraj, S.; Djukanovic, V.; Gasior, S.; Silanskas, A.; Venclovas, Č.; Siksnys, V. PAM Recognition by Miniature CRISPR–Cas12f Nucleases Triggers Programmable Double-Stranded DNA Target Cleavage. Nucleic Acids Res. 2020, 48 (9), 5016–5023. https://doi.org/10.1093/nar/gkaa208.
(21) Özcan, A.; Krajeski, R.; Ioannidi, E.; Lee, B.; Gardner, A.; Makarova, K. S.; Koonin, E. V; Abudayyeh, O. O.; Gootenberg, J. S. Programmable RNA Targeting with the Single-Protein CRISPR Effector Cas7-11. Nature 2021, 597 (7878), 720–725. https://doi.org/10.1038/s41586-021-03886-5.
Great Post! Keep up the good work! Marketplace Icon
LikeLike