PDF version: Global Highlights in Neuroengineering August 2005 to April 2019 – Logan Thrasher Collins
Optogenetic stimulation using ChR2: August 2005
Reference: (Boyden, Zhang, Bamberg, Nagel, & Deisseroth, 2005)
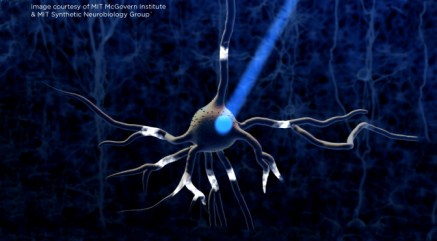
- Ed Boyden, Karl Deisseroth, and colleagues developed optogenetics, a revolutionary technique for stimulating neural activity.
- Optogenetics involves engineering neurons to express light-gated ion channels. The first channel used for this purpose was ChR2 (a protein originally found in bacteria which responds to blue light). In this way, a neuron exposed to an appropriate wavelength of light will be stimulated.
- Over time, optogenetics has gained a place as an essential experimental tool for neuroscientists across the world. It has been expanded upon and improved in numerous ways and has even allowed control of animal behavior via implanted fiber optics and other light sources. Optogenetics may eventually be used in the development of improved brain-computer interfaces.
Plans for Blue Brain Project announced
Reference: (Markram, 2006)
- Henry Markram announced the European Blue Brain Project (which later gave rise to the Human Brain Project), a large-scale computational neuroscience initiative with the goal of simulating the brain in a supercomputer.
- Plans for the early stages of the Blue Brain Project included reconstructing the morphologies of neuronal cell types from the rat neocortex, experimentally characterizing their electrophysiological parameters, building a virtual neocortical column with ~10,000 multicompartmental Hodgkin-Huxley-type neurons and over ten million synapses, and running simulations of the virtual neocortical column on the Blue Gene/L supercomputer.
- These goals were met by the November of 2007 and the general properties of the simulations reflected the biological reality.
- It should be noted that the virtual column’s connectivity was defined according the patterns of connectivity found in biological rats, but that this involved the numbers of inputs and outputs quantified for given cell types rather than exact wiring. Furthermore, the spatial distributions of boutons forming synaptic terminals upon target cells were also did not mirror biology exactly, but still reflected biological averages.

Optogenetic silencing using halorhodopsin: March 2007
Reference: (Han & Boyden, 2007)
- Ed Boyden continued developing optogenetic tools to manipulate neural activity. Along with Xue Han, he expressed a codon-optimized version of a bacterial halorhodopsin (along with the ChR2 protein) in neurons.
- Upon exposure to yellow light, halorhodopsin pumps chloride ions into the cell, hyperpolarizing the membrane and inhibiting neural activity.
- Using halorhodopsin and ChR2, neurons could be easily activated and inhibited using yellow and blue light respectively.

Brainbow: November 2007
Reference: (Livet et al., 2007)
- Lichtman and colleagues used Cre/Lox recombination tools to create genes which express a randomized set of three or more differently-colored fluorescent proteins (XFPs) in a given neuron, labeling the neuron with a unique combination of colors. About ninety distinct colors were emitted across a population of genetically modified neurons.
- The detailed structures within neural tissue equipped with the Brainbow system can be imaged much more easily since neurons can be distinguished via color contrast.
- As a proof-of-concept, hundreds of synaptic contacts and axonal processes were reconstructed in a selected volume of the cerebellum. Several other neural structures were also imaged using Brainbow.
- The fluorescent proteins expressed by the Brainbow system are usable in vivo.

Anatomically simplified simulation of 22 million neurons
Reference: (Djurfeldt et al., 2008)
- As part of the European FACETs project, a simulation consisting of 22 million neurons and 11 billion synapses was carried out on the IBM Blue Gene/L supercomputer.
- This simulation incorporated Hodgkin-Huxley-type models of three cell types including excitatory pyramidal cells, inhibitory basket cells, and inhibitory RNSP cells. Pyramidal cells were multicompartmental and consisted of six compartments.
- The model organized its virtual cells into hypercolumns, each containing 100 minicolumns. These columns were intended to reflect the rough biological organization of cells found in layers II/III of the cortex. Neuronal connectivity came from a simple set of rules. This involved rules for each cell type making synapses on a specified set of other cell types (including some synapses between cells of the same type) and rules for projections within and between minicolumns.
- Using this setup, the virtual neurons were able to store memories in the form of patterns of population activity. As with the situation in biological brains, stimulation of a subset of the population activity responsible for a certain memory caused the rest of that pattern of population activity to emerge.
- The simulation also demonstrated several other emergent phenomena that occur in biological brains including sustained elevation of pyramidal cell membrane potential during periods of high population activity as well as an exponentially distributed inter-spike intervals during periods of high population activity.

High temporal precision optogenetics: January 2010
Reference: (Gunaydin et al., 2010)
- Karl Deisseroth, Peter Hegemann, and colleagues used protein engineering to improve the temporal resolution of optogenetic stimulation.
- Glutamic acid at position 123 in ChR2 was mutated to threonine, producing a new ion channel protein (dubbed ChETA).
- The ChETA protein allows for induction of spike trains with frequencies up to 200 Hz and greatly decreases the incidence of unintended spikes. Furthermore, ChETA eliminates plateau potentials (a phenomenon which interferes with precise control of neural activity).

Hardware for faster than real-time neuroscience simulations: May 2010
Reference: (Schemmel et al., 2010)
- Scientists working under the European FACETs project developed a neuromorphic hardware system called BrainScaleS with the ability to simulate networks of adaptive exponential integrate-and-fire neurons 1,000 to 100,000 times faster than biological real-time.
- Each neuromorphic wafer was able to simulate up to 180,000 neurons and 4•107 Furthermore, the design allowed for interconnection of multiple wafers in order to simulate larger neuronal populations. Each artificial neuron was able to receive up to 14,336 presynaptic inputs (a biologically realistic value).
- To facilitate computational neuroscience applications using the neuromorphic wafers, a Python-based software interface called PyNN was created. In this way, the hardware could more easily be configured to simulate a desired neuronal system.

Hippocampal prosthesis in rats: March 2012
Reference: (Berger et al., 2012)
- Theodore Berger and his team developed an artificial replacement for neurons which transmit information from the CA3 region to the CA1 region of the hippocampus.
- This cognitive prosthesis employs recording and stimulation electrodes along with a multi-input multi-output (MIMO) model to encode the information in CA3 and transfer it to CA1.
- The hippocampal prosthesis was shown to restore and enhance memory in rats as evaluated by behavioral testing and brain imaging.
In vivo superresolution microscopy for neuroimaging: February 2012
Reference: (Berning, Willig, Steffens, Dibaj, & Hell, 2012)
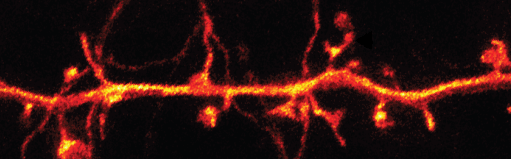
- Stefan Hell (2014 Nobel laureate in chemistry) developed stimulated emission depletion microscopy (STED), a type of superresolution fluorescence microscopy which allows imaging of synapses and dendritic spines.
- STED microscopy uses a torus-shaped de-excitation laser that interferes with the excitation laser to deplete fluorescence except in a very small spot. In this way, the diffraction limit is surpassed since the resulting light illuminates extremely small regions of the sample.
- Neurons in transgenic mice (equipped with glass-sealed holes in their skulls) were imaged using STED. Synapses and dendritic spines were observed up to fifteen nanometers below the surface of the brain tissue.
Eyewire as a crowdsourcing method for retina mapping: December 2012
Reference: (Marx, 2013)
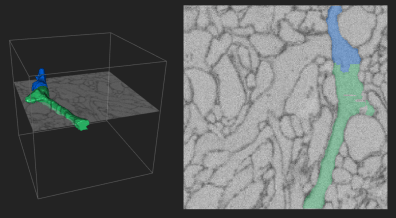
- The Eyewire project was created by Sebastian Seung’s research group. It is a crowdsourcing initiative for connectomic mapping within the retina towards uncovering neural circuits involved in visual processing.
- Laboratories first collect data via serial electron microscopy as well as functional data from two-photon microscopy.
- In the Eyewire game, images of tissue slices are provided to players who then help reconstruct neural morphologies and circuits by “coloring in” the parts of the images which correspond to cells and stacking many images on top of each other to generate 3D maps. Artificial intelligence tools help provide initial “best guesses” and guide the players, but the people ultimately perform the task of reconstruction.
- By November 2013, around 82,000 participants had played the game. Its popularity continues to grow.

In vivo three-photon microscopy: January 2013
Reference: (Horton et al., 2013)
- Multi-photon excitation uses pulsed lasers to excite fluorophores with two or more photons of light with long wavelengths. During the excitation, the photons undergo a nonlinear recombination process, yielding a single emitted photon with a much shorter wavelength. Because the excitation photons possess long wavelengths, they can penetrate tissue much more deeply than traditional microscopy allows.
- Horton and colleagues developed a three-photon excitation method to facilitate even deeper tissue penetration than the commonly used two-photon microscopic techniques.
- Since three photons were involved per excitation event, even longer excitation wavelengths (about 1,700 nm) were usable, allowing the construction of a 3-dimensional image stack that reached a depth of up to 1.4 mm within the living mouse brain.
- Blood vessels and RFP-labeled neurons were imaged using this approach. Furthermore, the depth was sufficient to enable imaging of neurons within the mouse hippocampus.

Whole-brain functional recording from larval zebrafish: March 2013
Reference: (Ahrens, Orger, Robson, Li, & Keller, 2013)
- Laser-scanning light-sheet microscopy was used to volumetrically image the entire brains of larval zebrafish (an optically transparent organism).
- The genetically encoded calcium sensor GCaMP5G facilitated functional recording at single-cell resolution from about 80% of the total neurons in the larval zebrafish brains. Computational methods were used to distinguish between individual neurons.
- Populations of neurons that underwent correlated activity patterns were identified to show the technique’s utility for uncovering the dynamics of neural circuits. These populations included hindbrain neurons that were functionally linked to neural activity in the spinal cord and a population of neurons which showed coupled oscillations on the left and right halves.

The BRAIN Initiative: April 2013
Reference: (“Fact Sheet: BRAIN Initiative,” 2013)
- The BRAIN Initiative (Brain Research through Advancing Innovative Technologies) provided neuroscientists with $110 million in governmental funding and $122 million in funding from private sources such as the Howard Hughes Medical Institute and the Allen Institute for Brain Science.
- The BRAIN Initiative focused on funding research which develops and utilizes new technologies for functional connectomics. It helped to accelerate research on tools for decoding the mechanisms of neural circuits in order to understand and treat mental illness, neurodegenerative diseases, and traumatic brain injury.
- The BRAIN Initiative emphasized collaboration between neuroscientists and physicists. It also pushed forward nanotechnology-based methods to image neural tissue, record from neurons, and otherwise collect neurobiological data.
The CLARITY method for making brains translucent: May 2013
Reference: (Chung & Deisseroth, 2013)
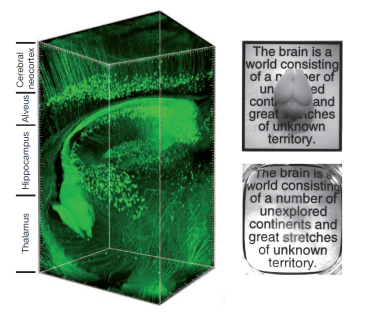
- Karl Deisseroth and colleagues developed a method called CLARITY to make samples of neural tissue optically translucent without damaging the fine cellular structures in the tissue. Using CLARITY, entire mouse brains have been turned transparent.
- Mouse brains were infused with hydrogel monomers (acrylamide and bisacrylamide) as well as formaldehyde and some other compounds for facilitating crosslinking. Next, the hydrogel monomers were crosslinked by incubating the brains at 37°C. Lipids in the hydrogel-stabilized mouse brains were extracted using hydrophobic organic solvents and electrophoresis.
- CLARITY allows antibody labeling, fluorescence microscopy, and other optically-dependent techniques to be used for imaging entire brains. In addition, it renders the tissue permeable to macromolecules, which broadens the types of experimental techniques that these samples can undergo (i.e. macromolecule-based stains, etc.)
X-ray microscopy reconstructs Drosophila brain hemisphere: September 2013
Reference: (Mizutani, Saiga, Takeuchi, Uesugi, & Suzuki, 2013)
- Mizutani and colleagues stained Drosophila brains with silver nitrate and tetrachloroaurate (a gold-containing compound), facilitating 3-dimensional imaging using X-ray microtomography at a voxel size of 220 × 328 × 314 nm.
- To generate the X-rays, a synchrotron source was used. It should be noted that synchrotron sources require large facilities to operate.
- Neuronal tracing was performed manually on the 3-dimensional X-ray images of the fly brain, a process which took about 1,700 person-hours. Some neuronal processes were too dense to be resolved, so they were “fused” into unified structures. Furthermore, some neuronal traces were fragmented and most of the cell bodies were not considered. This decreased the number of traces to one third of the estimated number of actual processes in the hemisphere.
- Mizutani’s investigation represents an early effort at large-scale connectomics that sets the stage for further initiatives as neuronal tracing, sample preparation, and X-ray microtomography technologies continue to improve.

Telepathic rats engineered using hippocampal prosthesis: December 2013
Reference: (Deadwyler et al., 2013)
- Berger’s hippocampal prosthesis was implanted in pairs of rats. When “donor” rats were trained to perform a task, they developed neural representations (memories) which were recorded by their hippocampal prostheses.
- The donor rat memories were run through the MIMO model and transmitted to the stimulation electrodes of the hippocampal prostheses implanted in untrained “recipient” rats. After receiving the memories, the recipient rats showed significant improvements on the task that they had not been trained to perform.

Integrated Information Theory 3.0: May 2014
Reference: (Oizumi, Albantakis, & Tononi, 2014)
- Integrated information theory (IIT) was originally proposed by Giulio Tononi in 2004. IIT is a quantitative theory of consciousness which may help explain the hard problem of consciousness.
- IIT begins by assuming the following phenomenological axioms; each experience is characterized by how it differs from other experiences, an experience cannot be reduced to interdependent parts, and the boundaries which distinguish individual experiences are describable as having defined “spatiotemporal grains.”
- From these phenomenological axioms and the assumption of causality, IIT identifies maximally irreducible conceptual structures (MICS) associated with individual experiences. MICS represent particular patterns of qualia that form unified percepts.
- IIT also outlines a mathematical measure of an experience’s quantity. This measure is called integrated information or ϕ.
Openworm: November 2014
Reference: (Szigeti et al., 2014)
- The anatomical elegans connectome was originally mapped in 1976 by Albertson and Thomson. More data has since been collected on neurotransmitters, electrophysiology, cell morphology, and other characteristics.
- Szigeti, Larson, and their colleagues made an online platform for crowdsourcing research on elegans computational neuroscience, with the goal of completing an entire “simulated worm.”
- The group also released software called Geppetto, a program that allows users to manipulate both multicompartmental Hodgkin-Huxley models and highly efficient soft-body physics simulations (for modeling the worm’s electrophysiology and anatomy).

Multibeam SEM for connectomics: January 2015
Reference: (Eberle et al., 2015)
- When mapping neuronal tissue with conventional scanning electron microscopy (SEM), the rate of data acquisition is severely limited by the amount of time it takes for the electron beam to image a single pixel on one slice of the sample. As such, imaging a 1 mm3 volume of tissue could take more than 25 years.
- By constructing a SEM device that produces a hexagonal array of 61 electron beams (as opposed to just one beam), Eberle and colleagues were able to greatly improve the throughput of SEM imaging. It was estimated that imaging a 1 mm3 volume with this system would take just 6 months.
- Multibeam SEM was designed for compatibility with both serial block-face techniques and automated serial sectioning methods.
- Slices of brain tissue, a bone tissue section, and a silicon wafer with printed circuitry were imaged with the multibeam SEM system as proof-of-concept tests.
Expansion microscopy: January 2015
Reference: (F. Chen, Tillberg, & Boyden, 2015)
- The Boyden group developed expansion microscopy, a method which enlarges neural tissue samples (including entire brains) with minimal structural distortions and so facilitates superior optical visualization of the scaled-up neural microanatomy. Furthermore, expansion microscopy greatly increases the optical translucency of treated samples.
- Expansion microscopy operates by infusing a swellable polymer network into brain tissue samples along with several chemical treatments to facilitate polymerization and crosslinking and then triggering expansion via dialysis in water. With 4.5-fold enlargement, expansion microscopy only distorts the tissue by about 1% (computed using a comparison between control superresolution microscopy of easily-resolvable cellular features and the expanded version).
- Before expansion, samples can express various fluorescent proteins to facilitate superresolution microscopy of the enlarged tissue once the process is complete. Furthermore, expanded tissue is highly amenable to fluorescent stains and antibody-based labels.

Neural lace: June 2015
Reference: (Liu et al., 2015)
- Charles Lieber’s group developed a syringe-injectable electronic mesh made of submicrometer-thick wiring for neural interfacing.
- The meshes were constructed using novel soft electronics for biocompatibility. Upon injection, the neural lace expands to cover and record from centimeter-scale regions of tissue.
- Neural lace may allow for “invasive” brain-computer interfaces to circumvent the need for surgical implantation. Lieber has continued to develop this technology towards clinical application.

BigNeuron towards standardized neuronal morphology acquisition: July 2015
Reference: (Peng et al., 2015)
- Because of the inconsistencies between neuronal reconstruction methods and lack of standardization found in neuronal morphology databases, BigNeuron was established as a community effort to improve the situation.
- BigNeuron tests as many automated neuronal reconstruction algorithms as possible using large-scale microscopy datasets (from several types of light microscopy). It uses the Vaa3D neuronal reconstruction software as a central platform. Reconstruction algorithms are added to Vaa3D as plugins. These computational tests are performed on supercomputers.
- BigNeuron aims to create a superior community-oriented neuronal morphology database, a set of greatly improved tools for neuronal reconstruction, a standardized protocol for future neuronal reconstructions, and a library of morphological feature definitions to facilitate classification.
The TrueNorth chip from DARPA and IBM: August 2015
Reference: (Akopyan et al., 2015)
- The TrueNorth neuromorphic computing chip was constructed and validated by DARPA and IBM. TrueNorth uses circuit modules which mimic neurons. Inputs to these fundamental circuit modules must overcome a threshold in order to trigger “firing.”
- The chip can emulate up to a million neurons with over 250 million synapses while requiring far less power than traditional computing devices.
Japan’s Brain/MINDS project: September 2015
Reference: (Okano, Miyawaki, & Kasai, 2015)
- In 2014, the Brain/MINDS (Brain Mapping by Integrated Neurotechnologies for Disease Studies) project was initiated to further neuroscientific understanding of the brain. This project received nearly $30 million in funding for its first year alone.
- Brain/MINDS focuses on studying the brain of the common marmoset (a non-human primate abundant in Japan), developing new technologies for brain mapping, and understanding the human brain with the goal of finding new treatments for brain diseases.
Human telepathy during a 20 questions game: September 2015
Reference: (Stocco et al., 2015)
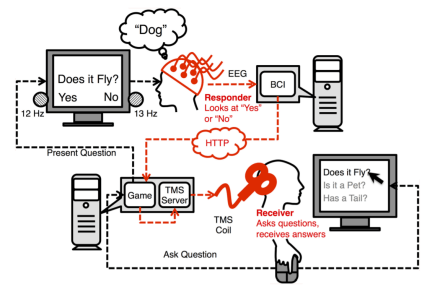
- Using an interactive question-and-answer setup, Stocco and colleagues demonstrated real-time telepathic communication between pairs of individuals via EEG and transcranial magnetic stimulation. Five pairs of participants played games of 20 questions and attempted to identify unknown objects.
- EEG data were recorded from the respondent, computationally processed, and transmitted as transcranial magnetic stimulation signals into the mind (occipital lobe stimulation) of a respondent. The respondent’s answers were translated into higher-intensity transcranial magnetic stimulation pulses corresponding to “yes” answers or lower-intensity transcranial magnetic stimulation pulses corresponding to “no” answers.
- When compared to control trials in which sham interfaces were used, the people using the brain-brain interfaces were significantly more successful at playing 20 questions games.
Human Brain Project cortical mesocircuit simulation: October 2015
Reference: (Markram et al., 2015)
- The Human Brain Project reconstructed a virtual version of a 0.29 mm3 region of rat cortical tissue including about 31,000 neurons and 37 million synapses.
- The connectivity was defined according the patterns of connectivity found in biological rats, (though this involved the numbers of inputs and outputs quantified for given cell types rather than explicit wiring). In addition, the spatial distributions of boutons forming synaptic terminals upon target cells reflected biological data.
- Detailed multicompartmental Hodgkin-Huxley-type models were used to take into account the effects of the morphologies of the neurons within the simulation.
- The cortical mesocircuit was emulated using the Blue Gene/Q supercomputer and this emulation was sufficiently accurate to reproduce emergent neurological processes and yield insights on the mechanisms of their computations.

Recording from C. elegans neurons reveals motor operations: October 2015
Reference: (Kato et al., 2015)
- Live elegans worms were immobilized in microfluidic devices and the neurons in their head ganglia as well as some of their motor systems were imaged and recorded from using the calcium indicator GCaMP. As the C. elegans connectome is well-characterized, Kato and colleagues were able to determine the identities of most of the cells that underwent imaging (with the help of computational segmentation techniques).
- Principal component analysis was used to reduce the dimensionality of the neural activity datasets since over 100 neurons per worm were recorded from simultaneously.
- Next, phase space analysis was utilized to visualize the patterns formed by the recording data. Motor behaviors including dorsal turns, ventral turns, forward movements, and backward movements were found to correspond to specific sequences of neural events as uncovered by examining the patterns found in the phase plots. Further analyses revealed various insights about these brain dynamics and their relationship to motor actions.

The China Brain Project: March 2016
Reference: (Poo et al., 2016)
- The China Brain Project was launched to help understand the neural mechanisms of cognition, develop brain research technology platforms, develop preventative and diagnostic interventions for brain disorders, and to improve brain-inspired artificial intelligence technologies.
- This project will take place from 2016 until 2030 with the goal of completing mesoscopic brain circuit maps.
- China’s population of non-human primates and preexisting non-human primate research facilities give the China Brain Project an advantage. The project will focus on studying rhesus macaques.
Expansion FISH: July 2016
Reference: (F. Chen et al., 2016)
- Boyden, Chen, Marblestone, Church, and colleagues combined fluorescent in situ hybridization (FISH) with expansion microscopy to image the spatial localization of RNA in neural tissue.
- The group developed a chemical linker to covalently attach intracellular RNA to the infused polymer network used in expansion microscopy. This allowed for RNAs to maintain their relative spatial locations within each cell post-expansion.
- After the tissue was enlarged, FISH was used to fluorescently label targeted RNA molecules. In this way, RNA localization was more effectively resolved.
- As a proof-of-concept, expansion FISH was used to reveal the nanoscale distribution of long noncoding RNAs in nuclei as well as the locations of RNAs within dendritic spines.
Neural dust: August 2016
Reference: (Seo et al., 2016)
- Michel Maharbiz’s group invented implantable, ~ 1 mm biosensors for wireless neural recording and tested them in rats.
- This neural dust could be miniaturized to less than 0.5 mm or even to microscale dimensions using customized electronic components.
- Neural dust motes consist of two recording electrodes, a transistor, and a piezoelectric crystal.
- The neural dust received external power from ultrasound. Neural signals were recorded by measuring disruptions to the piezoelectric crystal’s reflection of the ultrasound waves. Signal processing mathematics allowed precise detection of activity.

Bryan Johnson launches Kernel: August 2016
Reference: (Regalado, 2017)
- Entrepreneur Bryan Johnson invested $100 million to start Kernel, a neurotechnology company.
- Kernel plans to develop implants that allow for recording and stimulation of large numbers of neurons at once. The company’s initial goal is to develop treatments for mental illnesses and neurodegenerative diseases. Its long-term goal is to enhance human intelligence.
- Kernel originally partnered with Theodore Berger and intended to utilize his hippocampal prosthesis. Unfortunately, Berger and Kernel parted ways after about six months because Berger’s vision was reportedly too long-range to support a financially viable company (at least for now).
- Kernel was originally a company called Kendall Research Systems. This company was started by a former member of the Boyden lab. In total, four members of Kernel’s team are former Boyden lab members.
Somatosensory cortex stimulation for spinal cord injuries: October 2016
Reference: (Flesher et al., 2016)
- Gaunt, Flesher, and colleagues found that microstimulation of the primary somatosensory cortex (S1) partially restored tactile sensations to a patient with a spinal cord injury.
- Electrode arrays were implanted into the S1 regions of a patient with a spinal cord injury. The array performed intracortical microstimulation over a period of six months.
- The patient reported locations and perceptual qualities of the sensations elicited by microstimulation. The patient did not experience pain or “pins and needles” from any of the stimulus trains. Overall, 93% of the stimulus trains were reported as “possibly natural.”
- Results from this study might be used to engineer upper-limb neuroprostheses which provide somatosensory feedback.

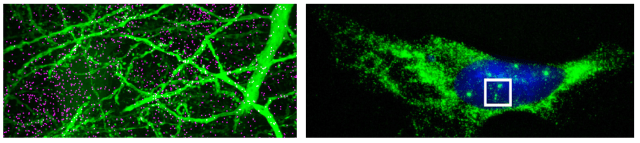
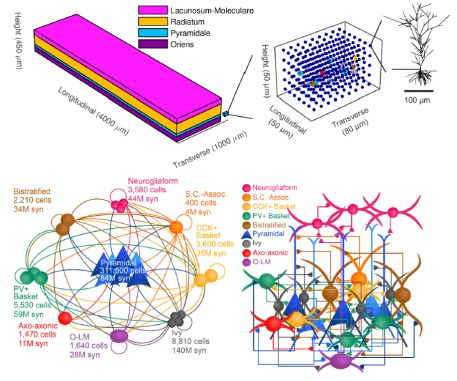
Simulation of rat CA1 region: December 2016
Reference: (Bezaire, Raikov, Burk, Vyas, & Soltesz, 2016)
- Detailed computational models of 338,740 neurons (including pyramidal cells and various types of interneurons) were equipped with connectivity patterns based on data from the biological CA1 region. External inputs were also estimated using biological data and incorporated into the simulation. It is important to note that these connectivity patterns described the typical convergence and divergence of neurites to and from particular cell types rather than explicitly representing the exact connections found in the biological rat.
- Each neuron was simulated using a multicompartmental Hodgkin-Huxley-type model with its morphological structure based on biological data from the given cell type. Furthermore, different cell types received different numbers of presynaptic terminals at specified distances from the soma. In total, over five billion synapses were present within the CA1 model.
- The simulation was implemented on several different supercomputers. Due to the model’s complexity, a four second simulation took about four hours to complete.
- As with the biological CA1 region, the simulation gave rise to gamma oscillations and theta oscillations as well as other biologically consistent phenomena. In addition, parvalbumin-expressing interneurons and neurogliaform cells were identified as drivers of the theta oscillations, demonstrating the utility of detailed neuronal simulations for uncovering biological insights.
The $100 billion Softbank Vision Fund: May 2017
Reference: (Lomas, 2017)
- Masayoshi Son, the CEO of Softbank (a Japanese telecommunications corporation), announced a plan to raise $100 billion in venture capital to invest in artificial intelligence. This plan involved partnering with multiple large companies in order to raise this enormous amount of capital.
- By the end of 2017, the Vision Fund successfully reached its $100 billion goal. Masayoshi Son has since announced further plans to continue raising money with a new goal of over $800 billion.
- Masayoshi Son’s reason for these massive investments is the technological singularity. He agrees with Kurzweil that the singularity will likely occur at around 2045 and he hopes to help bring the singularity to fruition. Though Son is aware of the risks posed by artificial superintelligence, he feels that superintelligent AI’s potential to tackle some of humanity’s greatest challenges (such as climate change and the threat of nuclear war) outweighs those risks.
UltraTracer enhances existing neuronal tracing software: March 2017
Reference: (Peng et al., 2017)
- UltraTracer is an algorithm that can improve the efficiency of existing neuronal tracing software for handling large datasets while maintaining accuracy.
- Datasets with hundreds of billions of voxels were utilized to test UltraTracer. Ten existing tracing algorithms were augmented.
- For most of the existing algorithms, the performance improvements were around 3-6 times, though a few showed improvements of 10-30 times. Even when using computers with smaller memory, UltraTracer was consistently able to enhance conventional software.
- UltraTracer was made opensource and is available as a plugin for the Vaa3D tracing software suite.
Elon Musk’s NeuraLink startup: March 2017
Reference: (Etherington, 2017)
- Elon Musk (CEO of Tesla, SpaceX, and a number of other successful companies) initiated a neuroengineering venture called NeuraLink.
- NeuraLink will begin by developing brain-computer interfaces (BCIs) for clinical applications, but the ultimate goal of the company is to enhance human cognitive abilities in order to keep up with artificial intelligence.
- Though many of the details around NeuraLink’s research are not yet open to the public, it has been rumored that injectable electronics similar to Lieber’s neural lace might be involved.
Facebook announces effort to build brain-computer interfaces: April 2017
Reference: (Constine, 2017)
- Facebook revealed research on constructing non-invasive brain-computer interfaces (BCIs) at a company-run conference in 2017. The initiative is run by Regina Dugan, Facebook’s head of R&D at division building 8.
- Facebook’s researchers are working on a non-invasive BCI which may eventually enable users to type one hundred words per minute with their thoughts alone. This effort builds on past investigations which have been used to help paralyzed patients.
- The building 8 group is also developing a wearable device for “skin hearing.” Using just a series of vibrating actuators which mimic the cochlea, test subjects have so far been able to recognize up to nine words. Facebook intends to vastly expand this device’s capabilities.
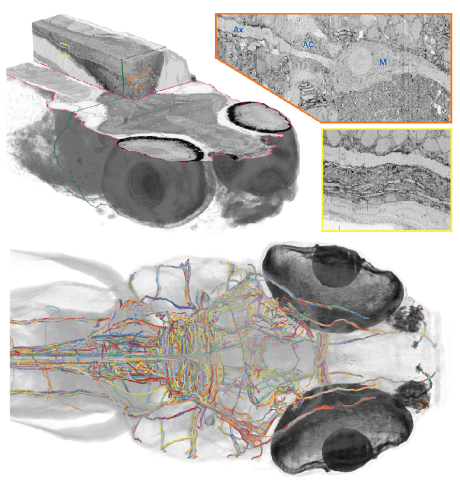
Whole-brain electron microscopy in larval zebrafish: May 2017
Reference: (Hildebrand et al., 2017)
- Serial electron microscopy facilitated imaging of the entire brain of a larval zebrafish at 5.5 days post-fertilization.
- Neuronal tracing software (a modified version of the CATMAID software) was used to reconstruct all the myelinated axons found in the larval zebrafish brain.
- The reconstructed dataset included 2,589 myelinated axon segments along with some of the associated soma and dendrites. It should be noted that only 834 of the myelinated axons were successfully traced back to their cell bodies.
Hardware for accelerated and biologically realistic simulations: May 2017
Reference: (Schemmel, Kriener, Müller, & Meier, 2017)
- Scientists working within the Human Brain Project developed an extension of the BrainScaleS neuromorphic hardware system for emulating biologically realistic neurons at a speed 1,000 times faster than biological real-time.
- The hardware’s design incorporates circuits blocks that act as compartments which represent segments of a dendritic tree. The organization and conductances of these compartments are configured using a system of switches. At a given time, each compartment can emulate either dendritic calcium spikes, dendritic NMDA spikes, or dendritic sodium spikes.
- Any electronic neuron within the chip can receive up to 16,000 presynaptic inputs as configured by the user. In addition, the presynaptic neurons feeding into a postsynaptic neuron can be located either on the same chip or on a different chip. In this way, combining multiple chips allows simulation of larger-scale systems.
- This neuromorphic system mimics biological plasticity using correlation sensors and a plasticity processing unit. The correlation sensors measure and locally store the exponentially weighted temporal difference between presynaptic and postsynaptic events within each electronic synapse. The plasticity processing unit reads these measurements along with the current weights and addresses of the synapses before implementing a software-defined algorithm for updating the synaptic weights and sometimes synaptic addresses. The algorithm can also be programmed to update the stored biophysical parameters of the circuit such as calcium spike threshold voltages, NMDA plateau durations, etc.
Human Brain Project analyzes brains using algebraic topology: June 2017
Reference: (Reimann et al., 2017)
- Investigators at the Human Brain Project utilized algebraic topology to analyze the reconstructed ~ 31,000 neuron cortical microcircuit from their earlier work.
- The analysis involved representing the cortical network as a digraph, finding directed cliques (complete directed subgraphs belonging to a digraph), and determining the net directionality of information flow (by computing the sum of the squares of the differences between in-degree and out-degree for all the neurons in a clique). In algebraic topology, directed cliques of n neurons are called directed simplices of dimension n-1.
- Vast numbers of high-dimensional directed cliques were found in the cortical microcircuit (as compared to null models and other controls). Spike correlations between pairs of neurons within a clique were found to increase with the clique’s dimension and with the proximity of the neurons to the clique’s sink. Furthermore, topological metrics allowed insights into the flow of neural information among multiple cliques.
- Experimental patch-clamp data supported the significance of the findings. In addition, similar patterns were found within the elegans connectome, suggesting that the results may generalize to nervous systems across species.

DARPA funds research to develop improved brain-computer interfaces: July 2017
Reference: (Hatmaker, 2017)
- The U.S. government agency DARPA awarded $65 million in total funding to six research groups.
- The recipients of this grant included five academic laboratories (headed by Arto Nurmikko, Ken Shepard, Jose-Alain Sahel and Serge Picaud, Vicent Pieribone, and Ehud Isacoff) and one small company called Paradromics Inc.
- DARPA’s goal for this initiative is to develop a nickel-sized bidirectional brain-computer interface (BCI) which can record from and stimulate up to one million individual neurons at once.
Early testing of hippocampal prosthesis algorithm in humans: July 2017
Reference: (Song, She, Hampson, Deadwyler, & Berger, 2017)
- Dong Song (who was working alongside Berger) tested the MIMO algorithm on human epilepsy patients using implanted recording and stimulation electrodes. The full hippocampal prosthesis was not implanted, but the electrodes acted similarly, though in a temporary capacity. Although only two patients were tested in this study, many trials were performed to compensate for the small sample size.
- Hippocampal spike trains from individual cells in CA1 and CA3 were recorded from the patients during a delayed match-to-sample task. The patients were shown various images while neural activity data were recorded by the electrodes and processed by the MIMO model. The patients were then asked to recall which image they had been shown previously by picking it from a group of “distractor” images. Memories encoded by the MIMO model were used to stimulate hippocampal cells during the recall phase.
- In comparison to controls in which the same two epilepsy patients were not assisted by the algorithm and stimulation, the experimental trials demonstrated a significant increase in successful pattern matching.
Brain imaging factory in China: August 2017
Reference: (Cyranoski, 2017)
- Qingming Luo started the HUST-Suzhou Institute for Brainsmatics, a brain imaging “factory.” Each of the numerous machines in Luo’s facility performs automated processing and imaging of tissue samples. The devices make ultrathin slices of brain tissue using diamond blades, treat the samples with fluorescent stains or other contrast-enhancing chemicals, and image then using fluorescence microscopy.
- The institute has already demonstrated its potential by mapping the morphology of a previously unknown neuron which “wraps around” the entire mouse brain.

Automated patch-clamp robot for in vivo neural recording: August 2017
Reference: (Suk et al., 2017)
- Ed Boyden and colleagues developed a robotic system to automate patch-clamp recordings from individual neurons. The robot was tested in vivo using mice and achieved a data collection yield similar to that of skilled human experimenters.
- By continuously imaging neural tissue using two-photon microscopy, the robot can adapt to a target cell’s movement and shift the pipette to compensate. This adaptation is facilitated by a novel algorithm called an imagepatching algorithm. As the pipette approaches its target, the algorithm adjusts the pipette’s trajectory based on the real-time two-photon microscopy.
- The robot can be used in vivo so long as the target cells express a fluorescent marker or otherwise fluoresce corresponding to their size and position.

Neuropixels probe: November 2017
Reference: (Jun et al., 2017)
- Jun and colleagues created the Neuropixels probe to facilitate simultaneous recording from hundreds of individual neurons with high spatiotemporal resolution. Previous extracellular probes were only able to record from a few dozen individual neurons.
- The Neuropixels recording shank is one centimeter long and includes 384 recording channels. Due to the small size of the accompanying apparatus (a 6×9 mm base and a data transmission cable), it enables high-throughput recording in freely moving animals. Because the shank is quite long, Neuropixels can record from multiple brain regions at once.
- Voltage signals are processed directly on the base of the Neuropixels apparatus, allowing for noise-free data transmission along the cable for further analysis.
![]()
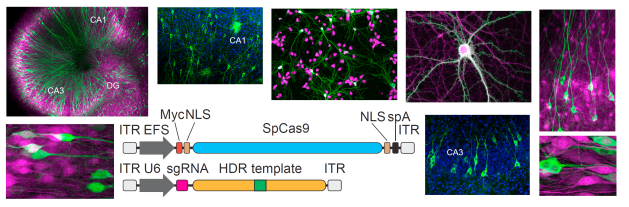
Genome editing in the mammalian brain: November 2017
Reference: (Nishiyama, Mikuni, & Yasuda, 2017)
- Precise genome editing in the brain has historically been challenging because most neurons are postmitotic (non-dividing) and the postmitotic state prevents homology-directed repair (HDR) from occurring. HDR is a mechanism of DNA repair which allows for targeted insertions of DNA fragments with overhangs homologous to the region of interest (by contrast, non-homologous end-joining is highly unpredictable).
- Nishiyama, Mikuni, and Yasuda developed a technique which allows genome editing in postmitotic mammalian neurons using adeno-associated viruses (AAVs) and CRISPR-Cas9.
- The AAVs delivered ssDNA sequences encoding a single guide RNA (sgRNA) and an insert. Inserts encoding a hemagglutinin tag (HA) and inserts encoding EGFP were both tested. Cas9 was encoded endogenously by transgenic host cells and in transgenic host animals.
- The technique achieved precise genome editing in vitro and in vivo with a low rate of off-target effects. Inserts did not cause deletion of nearby endogenous sequences for 98.1% of infected neurons.
SpiNNaker neuromorphic supercomputer
Reference: (Brown, Chad, Kamarudin, Dugan, & Furber, 2018)
- The SpiNNaker engine is a supercomputer designed to simulate spiking networks of neurons in real time. Its neuromorphic architecture facilitates optimal performance on these kinds of computational neuroscience tasks. SpiNNaker’s construction was funded by the Human Brain Project.
- SpiNNaker consists of a 240×240 mesh of nodes. Each node contains 18 ARM9 cores. Although ARM9 is a type of processor found in conventional computers, the communication infrastructure is less traditional. Only fixed-size packets of information can be transmitted among the cores. These packets are small, designed to carry the equivalent of a neuronal spike spike. Using an entirely hardware-based routing infrastructure, the information packets facilitate rapid communication throughout the machine.
- The system can be programmed to emulate any configuration of up to one billion neurons. It should be noted that this figure refers to point neuron models such as the leaky integrate-and-fire or Izhikevich models. However, it may also be possible to run multicompartmental models on SpiNNaker, though this would take more of its resources per neuron.

EEG-based facial image reconstruction: January 2018
Reference: (Nemrodov, Niemeier, Patel, & Nestor, 2018)
- EEG data associated with viewing images of faces was collected and used to determine the neural correlates of facial processing. In this way, the images were computationally reconstructed in a fashion resembling “mind reading.”
- It should be noted that the images reconstructed using data taken from multiple people were more accurate than the images reconstructed using single individuals. Nonetheless, the single individual data still yielded statistically significant accuracy.
- In addition to reconstructing the images themselves, the process gave insights on the cognitive steps involved in perceiving faces.

Upconversion nanoparticles for optogenetic stimulation: February 2018
Reference: (S. Chen et al., 2018)
- Upconversion nanoparticles absorb two or more low-energy photons and emit a higher energy photon. For instance, multiple near-infrared photons can be converted into a single visible spectrum photon.
- Shuo Chen and colleagues injected upconversion nanoparticles into the brains of mice and used them to convert externally applied near-infrared (NIR) light into visible light within the brain tissue. In this way, optogenetic stimulation was performed without the need for surgical implantation of fiber optics or similarly invasive procedures.
- The authors demonstrated stimulation via upconversion of NIR to blue light (to activate ChR2) and inhibition via upconversion of NIR to green light (to activate a rhodopsin called Arch).
- As a proof-of-concept, this technology was used to alter the behavior of the mice by activating hippocampally-encoded fear memories.

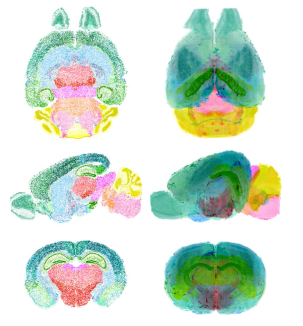
Map of all neuronal cell bodies within mouse brain: March 2018
Reference: (Murakami et al., 2018)
- Ueda, Murakami, and colleagues combined methods from expansion microscopy and CLARITY to develop a protocol called CUBIC-X which both expands and clears entire brains. Light-sheet fluorescence microscopy was used to image the treated brains and a novel algorithm was developed to detect individual nuclei.
- Although expansion microscopy causes some increased tissue transparency on its own, CUBIC-X greatly improved this property in the enlarged tissues, facilitating more detailed whole-brain imaging.
- Using CUBIC-X, the spatial locations of all the cell bodies (but not dendrites, axons, or synapses) within the mouse brain were mapped. This process was performed upon several adult mouse brains as well as several developing mouse brains to allow for comparative analysis.
- The authors made the spatial atlas publicly available in order to facilitate global cooperation towards annotating connectivity among the neural cell bodies within the atlas.
Clinical testing of hippocampal prosthesis algorithm in humans: March 2018
Reference: (Hampson et al., 2018)
- Further clinical tests of Berger’s hippocampal prosthesis were performed. Twenty-one patients took part in the experiments. Seventeen patients underwent CA3 recording so as to facilitate training and optimization of the MIMO model. Eight patients received CA1 stimulation so as to improve their memories.
- Electrodes with the ability to record from single neurons (10-24 single-neuron recording sites) and via EEG (4-6 EEG recording sites) were implanted such that recording and stimulation could occur at CA3 and CA1 respectively.
- Patients performed behavioral memory tasks. Both short-term and long-term memory showed an average improvement of 35% across the patients who underwent stimulation.
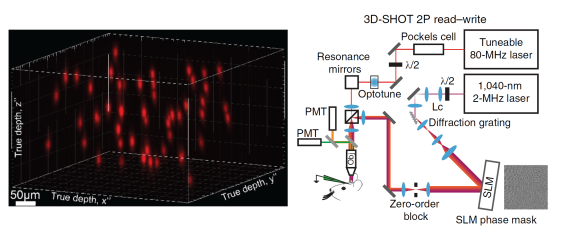
Precise optogenetic manipulation of fifty neurons: April 2018
Reference: (Mardinly et al., 2018)
- Mardinly and colleagues engineered a novel excitatory optogenetic ion channel called ST-ChroME and a novel inhibitory optogenetic ion channel called IRES-ST-eGtACR1. The channels were localized to the somas of host neurons and generated stronger photocurrents over shorter timescales than previously existing opsins, allowing for powerful and precise optogenetic stimulation and inhibition.
- 3D-SHOT is an optical technique in which light is tuned by a device called a spatial light modulator along with several other optical components. Using 3D-SHOT, light was precisely projected upon targeted neurons within a volume of 550×550×100 μm3.
- By combining novel optogenetic ion channels and the 3D-SHOT technique, complex patterns of neural activity were created in vivo with high spatial and temporal precision.
- Simultaneously, calcium imaging allowed measurement of the induced neural activity. More custom optoelectronic components helped avoid optical crosstalk of the fluorescent calcium markers with the photostimulating laser.
Whole-brain Drosophila image data via serial electron microscopy: July 2018
Reference: (Zheng et al., 2018)
- Zheng, Bock, and colleagues collected serial section electron microscopy data on the entire adult Drosophila brain, providing the data necessary to reconstruct a complete structural map of the fly’s brain at the resolution of individual synapses, dendritic spines, and axonal processes.
- The data are in the form of 7050 transmission electron microscopy images (187500 x 87500 pixels and 16 GB per image), each representing a 40nm-thin slice of the fly’s brain. In total the dataset requires 106 TB of storage.
- Although much of the data still needed to be processed to reconstruct a 3-dimensional map of the Drosophila brain, the authors did create 3-dimensional reconstructions of selected areas in the olfactory pathway of the fly. In doing so, they discovered a new cell type as well as several other previously unrealized insights about the organization of Drosophila‘s olfactory biology.

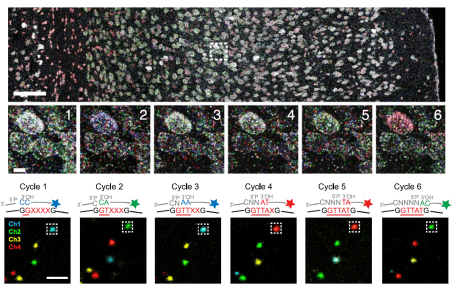
RNA-seq within intact brain tissue: July 2018
Reference: (Wang et al., 2018)
- Wang and colleagues developed a technique called STARmap for sequencing RNA within intact brain tissue, facilitating acquisition of three-dimensional spatial transcriptomic information.
- Fixed tissue was infused with a hydrogel matrix. DNA amplicons were covalently linked to the matrix via amine-modified nucleotides. Along with reverse transcriptase, these amplicons facilitated amplification of local mRNAs via rolling circle replication. Each amplicon contained a gene-unique barcode sequence to facilitate identification of genes. In addition, the tissue’s transparency was enhanced by lipid removal and protease treatment.
- To sequence the amplified mRNAs, two types of DNA probes were repeatedly washed through the tissue. Reading probes set the start position for decoding probes which were color-coded with fluorophores corresponding to the dinucleotides at their 3’ ends. With each cycle, the length of the reading probe was increased by one degenerate nucleotide, moving the reading start position ahead by one base. By observing the colors of the spots within the sample, each cycle facilitated identification of one nucleotide in all amplified mRNA molecules. After six or seven cycles of sequencing, the gene-unique barcode sequence associated with every mRNA could be identified. This sequencing process was called SEDAL.
- STARmap was used to quantify the expression of 160 genes in a thin slice of mouse cortex. These data reproduced the spatial gene expression pattern from a known dataset, validating STARmap.
- In thicker cortical volumes, the authors used STARmap to quantify spatial gene expression of 1020 genes across more than 30,000 cells. It was found that this number of genes represented an upper limit since trying to perform STARmap on a larger number of genes would result in too much overlap between the fluorescent spots.
Human telepathy using BrainNet: September 2018
Reference: (Jiang et al., 2018)
- EEG recordings were taken from two individuals (termed senders) while they played a Tetris-like game. Next, the recordings were converted into transcranial magnetic stimulation signals that acted to provide a third individual (called a receiver) with the necessary information to make decisions in the game without seeing the screen. The occipital cortex was stimulated. Fifteen people (five groups of three) took part in the study.
- To convey their information, the senders were told to focus upon either a higher or a lower intensity light corresponding to commands within the game (the two lights were placed on different sides of the computer screen). In the receiver’s mind, this translated to perceiving a flash of light. The receiver was able to distinguish the intensities and implement the correct command within the game.
- Using only the telepathically provided stimulation, the receiver made the correct game-playing decisions 81% of the time.

Transcriptomic cell type classification across mouse neocortex: October 2018
Reference: (Tasic et al., 2018)
- Single-cell RNA sequencing was used to characterize gene expression across 23,822 cells from the primary visual cortex and the anterior lateral motor cortex of mice.
- Using dimensionality reduction and clustering methods, the resulting data were used to classify the neurons into 133 transcriptomic cell types.
- Injections of adeno associated viruses (engineered to express fluorescent markers) facilitated retrograde tracing of neuronal projections within a subset of the sequenced cells. In this way, correspondences between projection patterns and transcriptomic identities were established.
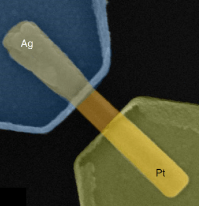
Nanowire system as a memristive artificial synapse: December 2018
Reference: (Milano et al., 2018)
- Milano and colleagues synthesized a memristive nanowire device consisting of a ZnO nanowire bridging between Ag and Pt electrodes on a SiO2 insulating substrate.
- When a positive voltage is applied to the Ag electrode, Ag+ ions start migrating along the nanowire towards the Pt electrode. Ag deposition on the nanowire creates a conductive bridge that allows for current to flow across the device. Each time voltage is applied, more Ag is deposited on the bridge, increasing the conductance across the terminals. In this way, the system mimics synaptic potentiation.
- Pulses with positive voltage across the terminals demonstrated the ability to increase the bridge’s conductance. Pulses with negative voltage were used to reset the nanowire bridge’s conductance back to the starting state.
- If shorter pulses are applied, the bridge’s conductance spontaneously decays back to its original state in a similar way to how biological synapses lose their potentiation after a period of inactivity.
- For a conductance increase to occur, a threshold amplitude of short pulses and a threshold number of consecutive short pulses were necessary. The amplitude and frequency of the pulses could be adjusted so that the kinetics of bridge formation and decay mimicked the kinetics of potentiation in biological synapses.
Whole-brain Drosophila image data acquired via expansion LLSM: January 2019
Reference: (Gao et al., 2019)
- Ed Boyden’s and Eric Betzig’s research groups combined lattice light-sheet microscopy with expansion microscopy to image the entire Drosophila brain as well as large volumes of mouse cortex at ~60×60×90 nm resolution. It should be noted that labeled subsets of structures were imaged rather than all structures within these volumes.
- In the Drosophila brain (a 340×660×90 μm volume), all ~110 dopaminergic neuronal membranes were immunostained with one color while all ~40 million presynaptic boutons were immunostained with a different color. Of these boutons, ~530,000 were associated with dopaminergic neurons. Eight of the dopaminergic cells underwent segmentation via manual tracing.
- Mouse somatosensory cortex (a ~100×150×150 μm subvolume) was imaged for the purpose of analyzing fluorescently labeled subcellular organelles within layer V pyramidal cells. 2893 mitochondria and 222 lysosomes were identified within five pyramidal cell bodies, five pyramidal cell apical dendrite initial portions, and three descending axon segments. These organelles were segmented and their morphological parameters were quantified.
- More mouse somatosensory cortex (a ~320×280×60 μm subvolume) was imaged with the purpose of examining cortical myelin sheaths. Myelin basic protein was immunostained in YFP-expressing pyramidal cells and the average structural properties of the myelin sheaths were quantified.
- Since the lattice light-sheet microscope had a limited 160 μm field of view, hundreds to thousands of tiles within the volumes were imaged and then computationally stitched together using customized software.

Pipeline for mesoscale mapping of the marmoset brain: February 2019
Reference: (Lin et al., 2019)
- Lin and colleagues established a histological pipeline for mapping mesoscale connectivity in the marmoset brain as part of Japan’s Brain/MINDS project. “Mesoscale” refers to connectivity between anatomical regions rather than individual neurons.
- MRI was performed upon living marmosets to determine proper locations for the injection of fluorescent neural tracers. Each marmoset brain was then sectioned into ~1700 slices with 20 μm thickness. In addition to the injected fluorescent tracers, the slices received a set of chemical stains to enhance visualization of neural features. The slices were then imaged.
- Using the images of the slices, three-dimensional reconstructions of marmoset brains were created. The MRI data helped to correct distortions in these reconstructions and to register the reconstructions into a standard three-dimensional atlas of the marmoset brain.
- The fluorescent tracer injections facilitated the construction of connectivity maps between distinct anatomical regions (i.e. V1 to V5, etc.) Projection strength between target and source regions was quantified using the number of voxels containing the fluorescent tracer labels.
- Using their experimental and computational pipeline, the authors predict that a mesoscale connectivity matrix for the entire marmoset brain could be finished by 2024 or earlier depending on the degree of collaboration with multiple groups.
Neural probes that resemble neurons: February 2019
Reference: (Yang et al., 2019)
- Charles Lieber’s group developed neural probes with the approximate size and shape of neurons (called NeuE probes). The NeuE probes consisted of soma-like platinum recording electrodes and neurite-like filaments made from a polymer insulator. At the core of the polymer, gold interconnect wiring carried current. The filaments were woven into a mesh.
- The electrodes exhibited similar dimensions compared with neuronal soma and the filaments had similar dimensions compared to dendrites and axons. In addition, the filaments demonstrated similar levels of flexibility relative to axons.
- The NeuE probes were highly biocompatible since they possessed similar size, shape, and flexibility compared to biological neurons. Microglia and astrocytes did not surround the probes, indicating a lack of immune response. Furthermore, neurons were not depleted around the probes. These results contrast with other neural implants that induce harmful immune reactions.
- NeuE probes with 16 electrodes were implanted in the hippocampal and cortical regions of living mouse brains. These probes demonstrated stable single-unit recording over a period of 90 days.
- After performing the recordings, the mice were killed and the brain tissue with the NeuE probes was imaged three-dimensionally using two-photon confocal microscopy. Mathematical analysis of the recording data helped identifiy which neurons were recorded by which electrodes.

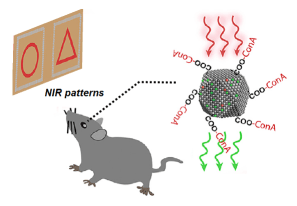
Injectable nanoparticles give mice infrared vision: April 2019
Reference: (Ma et al., 2019)
- Sub-retinal injections of upconversion nanoparticles gave mice the ability to perceive complex shapes on the near-infrared spectrum.
- The upconversion nanoparticles were linked to a protein that binds carbohydrates on the cell surface of photoreceptors. Since upconversion nanoparticles absorb longer wavelengths and then emit shorter wavelengths (i.e. near-infrared wavelengths are transformed to visible wavelengths), this allowed the mice to perceive infrared light patterns as if they were visible light.
- Treated mice retained their ability to see visible light alongside their new ability to see infrared light. The nanoparticle treatment also did not cause any adverse effects in the mice beyond some minor side effects (cataracts and corneal opacity) that happen with any sub-retinal injection. These minor side effects disappeared completely after two weeks. Furthermore, the mice still could distinguish complex near-infrared shapes ten weeks after injection of the nanoparticles, demonstrating the treatment’s long-term stability.
Reconstruction from whole-brain Drosophila image data
Reference: (Li et al., 2019)
- The whole-brain Drosophila electron microscopy data acquired by Zheng et al. underwent an automated image segmentation process to reconstruct all of the neurons in the fly’s brain. An algorithmic pipeline that involved flood-filling networks was used.
- It should be noted that the reconstruction is an “over-segmentation.” That is, each neuron is split into separate segments with an average length of 54 μm. However, improvements to the flood-filling algorithm may ameliorate this issue in the future. Furthermore, the separate segments can be manually joined at a rate ten times faster than full manual tracing of the neurons.

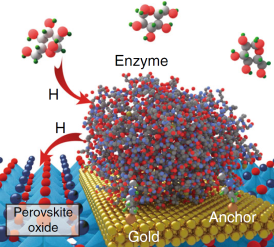
Perovskite nickelates directly interface with neurons: April 2019
Reference: (Zhang et al., 2019)
- A perovskite nickelate lattice with covalently linked horseradish peroxidase enzymes facilitated measurement of dopamine release from synapses within a mouse brain slice.
- When the dopaminergic neurons in the slice were stimulated, dopamine release triggered a hydrogen transfer process onto the perovskite nickelate lattice. This caused an increase in the lattice’s resistivity corresponding to the increase in dopamine concentration.
- Even in the complex media of an artificial cerebrospinal fluid, the device specifically detected dopamine. Furthermore, the device’s sensitivity was high enough that dopamine could be detected at concentrations as low as 5•10-17
- In separate experiments, a glucose oxidase enzyme was covalently linked to the perovskite nickelate lattice. When this device was exposed to glucose, hydrogen was transferred onto the lattice, causing a resistivity increase corresponding to the glucose concentration. The device only reacted with glucose and did not respond to mannose or galactose solutions. In addition, the device exhibited sufficient sensitivity to detect glucose at concentrations as low as 5•10-16
Decoding sentences from brain activity: April 2019
Reference: (Anumanchipalli, Chartier, & Chang, 2019)
- Anumanchipalli and colleagues constructed a system for translating neural activity into sentences.
- Electrocorticography (ECoG) was used to record the brain activity of several participating subjects while they spoke sentences aloud.
- A machine learning tool was used on audio recording data to create a model for inferring the facial movements that take place during speech. Another form of machine learning was used to predict inferred facial movements from brain activity. In addition, acoustic features were extracted from audio recordings of speech.
- By using the training data, the researchers developed a machine learning algorithm for mapping ECoG signals to inferred facial movements and inferred facial movements to sound.
- The algorithm allowed for generation of synthesized speech from brain activity. Listeners could often transcribe the synthesized sentences perfectly. However, some errors did occur. Nonetheless, the algorithm performed well enough that it could be very helpful to patients with disorders which prevent them from speaking.
References
Ahrens, M. B., Orger, M. B., Robson, D. N., Li, J. M., & Keller, P. J. (2013). Whole-brain functional imaging at cellular resolution using light-sheet microscopy. Nature Methods, 10, 413. Retrieved from https://doi.org/10.1038/nmeth.2434
Akopyan, F., Sawada, J., Cassidy, A., Alvarez-Icaza, R., Arthur, J., Merolla, P., … Modha, D. S. (2015). TrueNorth: Design and Tool Flow of a 65 mW 1 Million Neuron Programmable Neurosynaptic Chip. IEEE Transactions on Computer-Aided Design of Integrated Circuits and Systems, 34(10), 1537–1557. https://doi.org/10.1109/TCAD.2015.2474396
Anumanchipalli, G. K., Chartier, J., & Chang, E. F. (2019). Speech synthesis from neural decoding of spoken sentences. Nature, 568(7753), 493–498. https://doi.org/10.1038/s41586-019-1119-1
Berger, T. W., Song, D., Chan, R. H. M., Marmarelis, V. Z., LaCoss, J., Wills, J., … Granacki, J. J. (2012). A Hippocampal Cognitive Prosthesis: Multi-Input, Multi-Output Nonlinear Modeling and VLSI Implementation. IEEE Transactions on Neural Systems and Rehabilitation Engineering, 20(2), 198–211. https://doi.org/10.1109/TNSRE.2012.2189133
Berning, S., Willig, K. I., Steffens, H., Dibaj, P., & Hell, S. W. (2012). Nanoscopy in a Living Mouse Brain. Science, 335(6068), 551 LP-551. Retrieved from http://science.sciencemag.org/content/335/6068/551.abstract
Bezaire, M. J., Raikov, I., Burk, K., Vyas, D., & Soltesz, I. (2016). Interneuronal mechanisms of hippocampal theta oscillations in a full-scale model of the rodent CA1 circuit. ELife, 5, e18566. https://doi.org/10.7554/eLife.18566
Boyden, E. S., Zhang, F., Bamberg, E., Nagel, G., & Deisseroth, K. (2005). Millisecond-timescale, genetically targeted optical control of neural activity. Nature Neuroscience, 8, 1263. Retrieved from http://dx.doi.org/10.1038/nn1525
Brown, A. D., Chad, J. E., Kamarudin, R., Dugan, K. J., & Furber, S. B. (2018). SpiNNaker: Event-Based Simulation—Quantitative Behavior. IEEE Transactions on Multi-Scale Computing Systems, 4(3), 450–462. https://doi.org/10.1109/TMSCS.2017.2748122
Chen, F., Tillberg, P. W., & Boyden, E. S. (2015). Expansion microscopy. Science, 347(6221), 543 LP-548. https://doi.org/10.1126/science.1260088
Chen, F., Wassie, A. T., Cote, A. J., Sinha, A., Alon, S., Asano, S., … Boyden, E. S. (2016). Nanoscale imaging of RNA with expansion microscopy. Nature Methods, 13, 679. Retrieved from http://dx.doi.org/10.1038/nmeth.3899
Chen, S., Weitemier, A. Z., Zeng, X., He, L., Wang, X., Tao, Y., … McHugh, T. J. (2018). Near-infrared deep brain stimulation via upconversion nanoparticle–mediated optogenetics. Science, 359(6376), 679 LP-684. Retrieved from http://science.sciencemag.org/content/359/6376/679.abstract
Chung, K., & Deisseroth, K. (2013). CLARITY for mapping the nervous system. Nature Methods, 10, 508. Retrieved from http://dx.doi.org/10.1038/nmeth.2481
Constine, J. (2017). Facebook is building brain-computer interfaces for typing and skin-hearing. TechCrunch. Retrieved from https://techcrunch.com/2017/04/19/facebook-brain-interface/
Cyranoski, D. (2017). China launches brain-imaging factory. Nature, 548(7667), 268–269. https://doi.org/10.1038/548268a
Deadwyler, S., Hampson, R., Sweat, A., Song, D., Chan, R., Opris, I., … Berger, T. (2013). Donor/recipient enhancement of memory in rat hippocampus. Frontiers in Systems Neuroscience. Retrieved from https://www.frontiersin.org/article/10.3389/fnsys.2013.00120
Djurfeldt, M., Lundqvist, M., Johansson, C., Rehn, M., Ekeberg, O., & Lansner, A. (2008). Brain-scale simulation of the neocortex on the IBM Blue Gene/L supercomputer. IBM Journal of Research and Development, 52(1.2), 31–41. https://doi.org/10.1147/rd.521.0031
Eberle, A. L., Mikula, S., Schalek, R., Lichtman, J., Tate, M. L. K., & Zeidler, D. (2015). High-resolution, high-throughput imaging with a multibeam scanning electron microscope. Journal of Microscopy, 259(2), 114–120. https://doi.org/10.1111/jmi.12224
Etherington, D. (2017). Elon Musk’s Neuralink wants to boost the brain to keep up with AI. TechCrunch. Retrieved from techcrunch.com/2017/03/27/elon-musks-neuralink-wants-to-boost-the-brain-to-keep-up-with-ai/
Fact Sheet: BRAIN Initiative. (2013). Retrieved from https://obamawhitehouse.archives.gov/the-press-office/2013/04/02/fact-sheet-brain-initiative
Flesher, S. N., Collinger, J. L., Foldes, S. T., Weiss, J. M., Downey, J. E., Tyler-Kabara, E. C., … Gaunt, R. A. (2016). Intracortical microstimulation of human somatosensory cortex. Science Translational Medicine. Retrieved from http://stm.sciencemag.org/content/early/2016/10/12/scitranslmed.aaf8083.abstract
Gao, R., Asano, S. M., Upadhyayula, S., Pisarev, I., Milkie, D. E., Liu, T.-L., … Betzig, E. (2019). Cortical column and whole-brain imaging with molecular contrast and nanoscale resolution. Science, 363(6424), eaau8302. https://doi.org/10.1126/science.aau8302
Gunaydin, L. A., Yizhar, O., Berndt, A., Sohal, V. S., Deisseroth, K., & Hegemann, P. (2010). Ultrafast optogenetic control. Nature Neuroscience, 13, 387. Retrieved from http://dx.doi.org/10.1038/nn.2495
Hampson, R. E., Song, D., Robinson, B. S., Fetterhoff, D., Dakos, A. S., Roeder, B. M., … Deadwyler, S. A. (2018). Developing a hippocampal neural prosthetic to facilitate human memory encoding and recall. Journal of Neural Engineering, 15(3), 36014. https://doi.org/10.1088/1741-2552/aaaed7
Han, X., & Boyden, E. S. (2007). Multiple-Color Optical Activation, Silencing, and Desynchronization of Neural Activity, with Single-Spike Temporal Resolution. PLOS ONE, 2(3), e299. Retrieved from https://doi.org/10.1371/journal.pone.0000299
Hatmaker, T. (2017). DARPA awards $65 million to develop the perfect, tiny two-way brain-computer interface. TechCrunch. Retrieved from techcrunch.com/2017/07/10/darpa-nesd-grants-paradromics/
Hildebrand, D. G. C., Cicconet, M., Torres, R. M., Choi, W., Quan, T. M., Moon, J., … Engert, F. (2017). Whole-brain serial-section electron microscopy in larval zebrafish. Nature, 545, 345. Retrieved from https://doi.org/10.1038/nature22356
Horton, N. G., Wang, K., Kobat, D., Clark, C. G., Wise, F. W., Schaffer, C. B., & Xu, C. (2013). In vivo three-photon microscopy of subcortical structures within an intact mouse brain. Nature Photonics, 7, 205. Retrieved from https://doi.org/10.1038/nphoton.2012.336
Jiang, L., Stocco, A., Losey, D. M., Abernethy, J. A., Prat, C. S., & Rao, R. P. N. (2018). BrainNet: a multi-person brain-to-brain interface for direct collaboration between brains. ArXiv Preprint ArXiv:1809.08632.
Jun, J. J., Steinmetz, N. A., Siegle, J. H., Denman, D. J., Bauza, M., Barbarits, B., … Harris, T. D. (2017). Fully integrated silicon probes for high-density recording of neural activity. Nature, 551, 232. Retrieved from https://doi.org/10.1038/nature24636
Kato, S., Kaplan, H. S., Schrödel, T., Skora, S., Lindsay, T. H., Yemini, E., … Zimmer, M. (2015). Global Brain Dynamics Embed the Motor Command Sequence of Caenorhabditis elegans. Cell, 163(3), 656–669. https://doi.org/10.1016/j.cell.2015.09.034
Li, P. H., Lindsey, L. F., Januszewski, M., Zheng, Z., Bates, A. S., Taisz, I., … Jain, V. (2019). Automated Reconstruction of a Serial-Section EM Drosophila Brain with Flood-Filling Networks and Local Realignment. BioRxiv, 605634. https://doi.org/10.1101/605634
Lin, M. K., Takahashi, Y. S., Huo, B.-X., Hanada, M., Nagashima, J., Hata, J., … Mitra, P. (2019). A high-throughput neurohistological pipeline for brain-wide mesoscale connectivity mapping of the common marmoset. ELife, 8, e40042. https://doi.org/10.7554/eLife.40042
Liu, J., Fu, T.-M., Cheng, Z., Hong, G., Zhou, T., Jin, L., … Lieber, C. M. (2015). Syringe-injectable electronics. Nature Nanotechnology, 10, 629. Retrieved from http://dx.doi.org/10.1038/nnano.2015.115
Livet, J., Weissman, T. A., Kang, H., Draft, R. W., Lu, J., Bennis, R. A., … Lichtman, J. W. (2007). Transgenic strategies for combinatorial expression of fluorescent proteins in the nervous system. Nature, 450, 56. Retrieved from http://dx.doi.org/10.1038/nature06293
Lomas, N. (2017). Superintelligent AI explains Softbank’s push to raise a $100BN Vision Fund. TechCrunch. Retrieved from https://techcrunch.com/2017/02/27/superintelligent-ai-explains-softbanks-push-to-raise-a-100bn-vision-fund/
Ma, Y., Bao, J., Zhang, Y., Li, Z., Zhou, X., Wan, C., … Xue, T. (2019). Mammalian Near-Infrared Image Vision through Injectable and Self-Powered Retinal Nanoantennae. Cell, 177(2), 243–255.e15. https://doi.org/https://doi.org/10.1016/j.cell.2019.01.038
Mardinly, A. R., Oldenburg, I. A., Pégard, N. C., Sridharan, S., Lyall, E. H., Chesnov, K., … Adesnik, H. (2018). Precise multimodal optical control of neural ensemble activity. Nature Neuroscience, 21(6), 881–893. https://doi.org/10.1038/s41593-018-0139-8
Markram, H. (2006). The Blue Brain Project. Nature Reviews Neuroscience, 7, 153. Retrieved from http://dx.doi.org/10.1038/nrn1848
Markram, H., Muller, E., Ramaswamy, S., Reimann, M. W., Abdellah, M., Sanchez, C. A., … Schürmann, F. (2015). Reconstruction and Simulation of Neocortical Microcircuitry. Cell, 163(2), 456–492. https://doi.org/10.1016/j.cell.2015.09.029
Marx, V. (2013). Neuroscience waves to the crowd. Nature Methods, 10, 1069. Retrieved from http://dx.doi.org/10.1038/nmeth.2695
Milano, G., Luebben, M., Ma, Z., Dunin-Borkowski, R., Boarino, L., Pirri, C. F., … Valov, I. (2018). Self-limited single nanowire systems combining all-in-one memristive and neuromorphic functionalities. Nature Communications, 9(1), 5151. https://doi.org/10.1038/s41467-018-07330-7
Mizutani, R., Saiga, R., Takeuchi, A., Uesugi, K., & Suzuki, Y. (2013). Three-dimensional network of Drosophila brain hemisphere. Journal of Structural Biology, 184(2), 271–279. https://doi.org/https://doi.org/10.1016/j.jsb.2013.08.012
Murakami, T. C., Mano, T., Saikawa, S., Horiguchi, S. A., Shigeta, D., Baba, K., … Ueda, H. R. (2018). A three-dimensional single-cell-resolution whole-brain atlas using CUBIC-X expansion microscopy and tissue clearing. Nature Neuroscience, 21(4), 625–637. https://doi.org/10.1038/s41593-018-0109-1
Nemrodov, D., Niemeier, M., Patel, A., & Nestor, A. (2018). The Neural Dynamics of Facial Identity Processing: Insights from EEG-Based Pattern Analysis and Image Reconstruction. Eneuro, 5(1), ENEURO.0358-17.2018. https://doi.org/10.1523/ENEURO.0358-17.2018
Nishiyama, J., Mikuni, T., & Yasuda, R. (2017). Virus-Mediated Genome Editing via Homology-Directed Repair in Mitotic and Postmitotic Cells in Mammalian Brain. Neuron, 96(4), 755–768.e5. https://doi.org/10.1016/j.neuron.2017.10.004
Oizumi, M., Albantakis, L., & Tononi, G. (2014). From the Phenomenology to the Mechanisms of Consciousness: Integrated Information Theory 3.0. PLOS Computational Biology, 10(5), e1003588. Retrieved from https://doi.org/10.1371/journal.pcbi.1003588
Okano, H., Miyawaki, A., & Kasai, K. (2015). Brain/MINDS: brain-mapping project in Japan. Philosophical Transactions of the Royal Society of London. Series B, Biological Sciences, 370(1668). https://doi.org/10.1098/rstb.2014.0310
Peng, H., Hawrylycz, M., Roskams, J., Hill, S., Spruston, N., Meijering, E., & Ascoli, G. A. (2015). BigNeuron: Large-Scale 3D Neuron Reconstruction from Optical Microscopy Images. Neuron, 87(2), 252–256. https://doi.org/10.1016/j.neuron.2015.06.036
Peng, H., Zhou, Z., Meijering, E., Zhao, T., Ascoli, G. A., & Hawrylycz, M. (2017). Automatic tracing of ultra-volumes of neuronal images. Nature Methods, 14, 332. Retrieved from https://doi.org/10.1038/nmeth.4233
Poo, M., Du, J., Ip, N. Y., Xiong, Z.-Q., Xu, B., & Tan, T. (2016). China Brain Project: Basic Neuroscience, Brain Diseases, and Brain-Inspired Computing. Neuron, 92(3), 591–596. https://doi.org/10.1016/j.neuron.2016.10.050
Regalado, A. (2017). The Entrepreneur with the $100 Million Plan to Link Brains to Computers. MIT Technology Review. Retrieved from https://www.technologyreview.com/s/603771/the-entrepreneur-with-the-100-million-plan-to-link-brains-to-computers/
Reimann, M. W., Nolte, M., Scolamiero, M., Turner, K., Perin, R., Chindemi, G., … Markram, H. (2017). Cliques of Neurons Bound into Cavities Provide a Missing Link between Structure and Function. Frontiers in Computational Neuroscience. Retrieved from https://www.frontiersin.org/article/10.3389/fncom.2017.00048
Schemmel, J., Briiderle, D., Griibl, A., Hock, M., Meier, K., & Millner, S. (2010). A wafer-scale neuromorphic hardware system for large-scale neural modeling. In Proceedings of 2010 IEEE International Symposium on Circuits and Systems (pp. 1947–1950). https://doi.org/10.1109/ISCAS.2010.5536970
Schemmel, J., Kriener, L., Müller, P., & Meier, K. (2017). An accelerated analog neuromorphic hardware system emulating NMDA- and calcium-based non-linear dendrites. In 2017 International Joint Conference on Neural Networks (IJCNN) (pp. 2217–2226). https://doi.org/10.1109/IJCNN.2017.7966124
Seo, D., Neely, R. M., Shen, K., Singhal, U., Alon, E., Rabaey, J. M., … Maharbiz, M. M. (2016). Wireless Recording in the Peripheral Nervous System with Ultrasonic Neural Dust. Neuron, 91(3), 529–539. https://doi.org/10.1016/j.neuron.2016.06.034
Song, D., She, X., Hampson, R. E., Deadwyler, S. A., & Berger, T. W. (2017). Multi-resolution multi-trial sparse classification model for decoding visual memories from hippocampal spikes in human. In 2017 39th Annual International Conference of the IEEE Engineering in Medicine and Biology Society (EMBC) (pp. 1046–1049). https://doi.org/10.1109/EMBC.2017.8037006
Stocco, A., Prat, C. S., Losey, D. M., Cronin, J. A., Wu, J., Abernethy, J. A., & Rao, R. P. N. (2015). Playing 20 Questions with the Mind: Collaborative Problem Solving by Humans Using a Brain-to-Brain Interface. PLOS ONE, 10(9), e0137303. Retrieved from https://doi.org/10.1371/journal.pone.0137303
Suk, H.-J., van Welie, I., Kodandaramaiah, S. B., Allen, B., Forest, C. R., & Boyden, E. S. (2017). Closed-Loop Real-Time Imaging Enables Fully Automated Cell-Targeted Patch-Clamp Neural Recording In Vivo. Neuron, 95(5), 1037–1047.e11. https://doi.org/https://doi.org/10.1016/j.neuron.2017.08.011
Szigeti, B., Gleeson, P., Vella, M., Khayrulin, S., Palyanov, A., Hokanson, J., … Larson, S. (2014). OpenWorm: an open-science approach to modeling Caenorhabditis elegans. Frontiers in Computational Neuroscience. Retrieved from https://www.frontiersin.org/article/10.3389/fncom.2014.00137
Tasic, B., Yao, Z., Graybuck, L. T., Smith, K. A., Nguyen, T. N., Bertagnolli, D., … Zeng, H. (2018). Shared and distinct transcriptomic cell types across neocortical areas. Nature, 563(7729), 72–78. https://doi.org/10.1038/s41586-018-0654-5
Wang, X., Allen, W. E., Wright, M. A., Sylwestrak, E. L., Samusik, N., Vesuna, S., … Deisseroth, K. (2018). Three-dimensional intact-tissue sequencing of single-cell transcriptional states. Science, 361(6400), eaat5691. https://doi.org/10.1126/science.aat5691
Yang, X., Zhou, T., Zwang, T. J., Hong, G., Zhao, Y., Viveros, R. D., … Lieber, C. M. (2019). Bioinspired neuron-like electronics. Nature Materials, 18(5), 510–517. https://doi.org/10.1038/s41563-019-0292-9
Zhang, H.-T., Zuo, F., Li, F., Chan, H., Wu, Q., Zhang, Z., … Ramanathan, S. (2019). Perovskite nickelates as bio-electronic interfaces. Nature Communications, 10(1), 1651. https://doi.org/10.1038/s41467-019-09660-6
Zheng, Z., Lauritzen, J. S., Perlman, E., Robinson, C. G., Nichols, M., Milkie, D., … Bock, D. D. (2018). A Complete Electron Microscopy Volume of the Brain of Adult Drosophila melanogaster. Cell, 174(3), 730–743.e22. https://doi.org/10.1016/j.cell.2018.06.019